Artificial intelligence and first-principle methods in protein redesign: A marriage of convenience?
- PMID: 40671352
- PMCID: PMC12267658
- DOI: 10.1002/pro.70210
Artificial intelligence and first-principle methods in protein redesign: A marriage of convenience?
Abstract
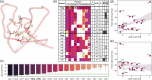
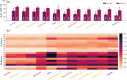
Since AlphaFold2's rise, many deep learning methods for protein design have emerged. Here, we validate widely used and recognized tools, compare them with first-principle methods, and explore their combinations, focusing on their effectiveness in protein redesign and potential for therapeutic repurposing. We address two challenges: evaluating tools and combinations ability to detect the effects of multiple concurrent mutations in protein variants, and leveraging large-scale datasets to compare modeling-free methods, namely force fields, which handle point mutations well with limited backbone rearrangement, and inverse folding tools, which excel at native sequence recovery but may struggle with non-natural proteins. Debuting TriCombine, a tool that identifies residue triangles in input structures, matches them to a structural database, and scores mutants based on substitution frequencies, we shortlisted candidates, modeled them with FoldX, and generated 16 SH3 mutants carrying up to 9 concurrent substitutions. The dataset was expanded to include 36 mutants and 11 crystal structures (7 newly solved), along with a parallel set of multiple non-concurrent mutants from three additional proteins. For broader validation, we analyzed 160,000 four-site GB1 mutants and 163,555 (single and double) variants across 179 natural and de novo domains. We show that combining AI-based modeling tools with force field scoring functions yields the most reliable results. Inverse folding tools perform very well but lose accuracy on less-represented proteins. First-principle force fields like FoldX remain the most accurate for point mutations. All methods perform worse when applied to unsolved de novo models, underscoring the need for hybrid strategies in robust protein design.
Keywords: artificial intelligence; crystallographic structure; force field; protein design.
© 2025 The Author(s). Protein Science published by Wiley Periodicals LLC on behalf of The Protein Society.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures


References
-
- Ahdritz G, Bouatta N, Kadyan S, Xia Q, Gerecke W, O'Donnell TJ, et al. OpenFold: Retraining AlphaFold2 yields new insights into its learning mechanisms and capacity for generalization. 2022. bioRxiv [Internet]. Available from: http://biorxiv.org/lookup/doi/10.1101/2022.11.20.517210 - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

