This is a preprint.
ENHANCED CLEAVAGE OF GENOMIC CCR5 USING CASX2Max
- PMID: 40672203
- PMCID: PMC12265720
- DOI: 10.1101/2025.07.08.663680
ENHANCED CLEAVAGE OF GENOMIC CCR5 USING CASX2Max
Abstract
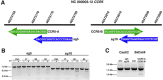
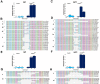
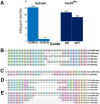
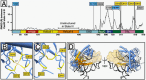
Development of novel CRISPR/Cas systems enhances opportunities for gene editing to treat infectious diseases, cancer, and genetic disorders. We evaluated CasX2 (PlmCas12e), a class II CRISPR system derived from Planctomycetes, a non-pathogenic bacterium present in aquatic and terrestrial soils. CasX2 offers several advantages over Streptococcus pyogenes Cas9 (SpCas9) and Staphylococcus aureus Cas9 (SaCas9), including its smaller size, distinct protospacer adjacent motif (PAM) requirements, staggered cleavage cuts that promote homology-directed repair, and no known pre-existing immunity in humans. A recent study reported that a three amino acid substitution in CasX2 significantly enhanced cleavage activity (1). Therefore, we compared cleavage efficiency and double-stranded break repair characteristics between the native CasX2 and the variant, CasX2Max, for cleavage of CCR5, a gene that encodes the CCR5 receptor important for HIV-1 infection. Two CasX2 single guide RNAs (sgRNAs) were designed that flanked the 32 bases deleted in the natural CCR5 Δ32 mutation. Nanopore sequencing demonstrated that CasX2 using sgRNAs with spacers of 17 nucleotides (nt), 20 nt or 23 nt in length were ineffective at cleaving genomic CCR5. In contrast, CasX2Max using sgRNAs with 20 nt and 23 nt spacer lengths, enabled robust genomic cleavage of CCR5. Structural modeling indicated that two of the CasX2Max substitutions enhanced sgRNA-DNA duplex stability, while the third improved DNA strand alignment within the catalytic site. These structural changes likely underlie the increased activity of CasX2Max in cellular gene excision. In sum, CasX2Max consistently outperformed native CasX2 across all assays and represents a superior gene-editing platform for therapeutic applications.
Keywords: CCR5; CRISPR/Cas; CasX2; guide RNA.
Figures





Similar articles
-
PlmCas12e (CasX2) cleavage of CCR5: impact of guide RNA spacer length and PAM sequence on cleavage activity.RNA Biol. 2023 Jan;20(1):296-305. doi: 10.1080/15476286.2023.2221510. RNA Biol. 2023. PMID: 37287312 Free PMC article.
-
CAS12e (CASX2) CLEAVAGE OF CCR5: IMPACT OF GUIDE RNA LENGTH AND PAM SEQUENCE ON CLEAVAGE ACTIVITY.bioRxiv [Preprint]. 2023 Jan 2:2023.01.02.522476. doi: 10.1101/2023.01.02.522476. bioRxiv. 2023. Update in: RNA Biol. 2023 Jan;20(1):296-305. doi: 10.1080/15476286.2023.2221510. PMID: 36711562 Free PMC article. Updated. Preprint.
-
[Improving the Efficiency and Safety of Human CCR5 Gene Editing by Selection of Optimal Guide RNAs for SpCAS9 and CAS12A].Mol Biol (Mosk). 2025 Mar-Apr;59(2):234-243. Mol Biol (Mosk). 2025. PMID: 40558035 Russian.
-
Antiretrovirals for reducing the risk of mother-to-child transmission of HIV infection.Cochrane Database Syst Rev. 2011 Jul 6;(7):CD003510. doi: 10.1002/14651858.CD003510.pub3. Cochrane Database Syst Rev. 2011. PMID: 21735394
-
Drugs for preventing postoperative nausea and vomiting in adults after general anaesthesia: a network meta-analysis.Cochrane Database Syst Rev. 2020 Oct 19;10(10):CD012859. doi: 10.1002/14651858.CD012859.pub2. Cochrane Database Syst Rev. 2020. PMID: 33075160 Free PMC article.
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
