Carboxylic Acid Bioisosteres Boost Nurr1 Agonist Selectivity
- PMID: 40674328
- PMCID: PMC7617969
- DOI: 10.1021/acs.jmedchem.5c01140
Carboxylic Acid Bioisosteres Boost Nurr1 Agonist Selectivity
Abstract
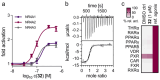
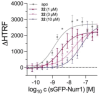
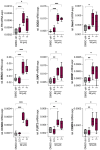
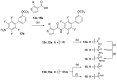
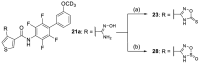
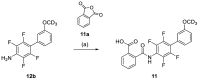
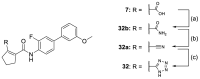
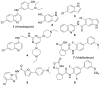
Nuclear receptor related 1 (Nurr1) is a neuronal ligand-activated transcription factor implicated in neurodegenerative diseases including Alzheimer's disease, Parkinson's disease and multiple sclerosis, which has fueled the development of Nurr1 modulators. Among them, the clinically studied dihydroorotate dehydrogenase (DHODH) inhibitor vidofludimus was found to exhibit strong Nurr1 agonism. Here, we aimed to establish a vidofludimus-derived Nurr1 agonist lacking DHODH inhibitor potency as a tool. We explored bioisosteric replacement of the drug's carboxylate motif and succeeded in boosting selectivity for Nurr1 over DHODH to >100-fold. Dopaminergic neural cells treated with the optimized tetrazole-based Nurr1 agonist revealed induction of genes involved in neuroprotection and neuronal health, supporting the potential of Nurr1 activation in neurodegenerative diseases.
Conflict of interest statement
CG, HK and DV hold patent application(s) claiming new compounds described in this study.
Figures
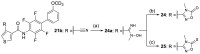
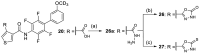
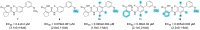
Similar articles
-
Targeting dihydroorotate dehydrogenase (DHODH) for host-directed therapy: Discovery of a novel alkyne-based scaffold inhibitor effective against SARS-CoV-2.Eur J Med Chem. 2025 Nov 5;297:117876. doi: 10.1016/j.ejmech.2025.117876. Epub 2025 Jun 21. Eur J Med Chem. 2025. PMID: 40592187
-
Direct long-read visualization reveals hidden variation in GCH1 gene copy number and precise expansion steps.BMC Genomics. 2025 Jul 17;26(1):671. doi: 10.1186/s12864-025-11859-5. BMC Genomics. 2025. PMID: 40676505 Free PMC article.
-
Development of a Potent Nurr1 Agonist Tool for In Vivo Applications.J Med Chem. 2023 May 11;66(9):6391-6402. doi: 10.1021/acs.jmedchem.3c00415. Epub 2023 Apr 26. J Med Chem. 2023. PMID: 37127285 Free PMC article.
-
DHODH-mediated mitochondrial redox homeostasis: a novel ferroptosis regulator and promising therapeutic target.Redox Biol. 2025 Sep;85:103788. doi: 10.1016/j.redox.2025.103788. Epub 2025 Jul 23. Redox Biol. 2025. PMID: 40716151 Free PMC article. Review.
-
Nurr1 modulators - a patent review (2019-present).Expert Opin Ther Pat. 2025 Aug;35(8):795-810. doi: 10.1080/13543776.2025.2516780. Epub 2025 Jun 18. Expert Opin Ther Pat. 2025. PMID: 40481732 Review.
References
-
- Wang Z, Benoit G, Liu J, Prasad S, Aarnisalo P, Liu X, Xu H, Walker NPC, Perlmann T. Structure and Function of Nurr1 Identifies a Class of Ligand- Independent Nuclear Receptors. Nature. 2003;423(6939):555–560. - PubMed
-
- Jakaria M, Haque ME, Cho D-Y, Azam S, Kim I-S, Choi D-K. Molecular Insights into NR4A2(Nurr1): An Emerging Target for Neuroprotective Therapy Against Neuroinflammation and Neuronal Cell Death. Mol Neurobiol. 2019;56(8):5799–5814. - PubMed
-
- Liu W, Gao Y, Chang N. Nurr1 Overexpression Exerts Neuroprotective and Anti-Inflammatory Roles via down-Regulating CCL2 Expression in Both in Vivo and in Vitro Parkinson’s Disease Models. Biochem Biophys Res Commun. 2017;482(4):1312–1319. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

