Baseline predictors for 28-day COVID-19 severity and mortality among hospitalized patients: results from the IMPACC study
- PMID: 40687705
- PMCID: PMC12271175
- DOI: 10.3389/fmed.2025.1604388
Baseline predictors for 28-day COVID-19 severity and mortality among hospitalized patients: results from the IMPACC study
Abstract
Introduction: The coronavirus disease 2019 (COVID-19) pandemic threatened public health and placed a significant burden on medical resources. The Immunophenotyping Assessment in a COVID-19 Cohort (IMPACC) study collected clinical, demographic, blood cytometry, serum receptor-binding domain (RBD) antibody titers, metabolomics, targeted proteomics, nasal metagenomics, Olink, nasal viral load, autoantibody, SARS-CoV-2 antibody titers, and nasal and peripheral blood mononuclear cell (PBMC) transcriptomics data from patients hospitalized with COVID-19. The aim of this study is to select baseline biomarkers and build predictive models for 28-day in-hospital COVID-19 severity and mortality with most predictive variables while prioritizing routinely collected variables.
Methods: We analyzed 1102 hospitalized COVID-19 participants. We used the lasso and forward selection to select top predictors for severity and mortality, and built predictive models based on balanced training data. We then validated the models on testing data.
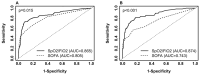
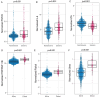
Results: Severity was best predicted by the baseline SpO2/FiO2 ratio obtained from COVID-19 patients (test AUC: 0.874). Adding patient age, BMI, FGF23, IL-6, and LTA to the disease severity prediction model improves the test AUC by an additional 3%. The clinical mortality prediction model using SpO2/FiO2 ratio, age, and BMI resulted in a test AUC of 0.83. Adding laboratory results such as TNFRSF11B and plasma ribitol count increased the prediction model by 3.5%. The severity and mortality prediction models developed outperform the Sequential Organ Failure Assessment (SOFA) score among inpatients and perform similarly to the SOFA score among ICU patients.
Conclusion: This study identifies clinical data and laboratory biomarkers of COVID-19 severity and mortality using machine learning models. The study identifies SpO2/FiO2 ratio to be the most important predictor for both severity and mortality. Several biomarkers were identified to modestly improve the predictions. The results also provide a baseline of SARS-CoV-2 infection during the early stages of the coronavirus emergence and can serve as a baseline for future studies that inform how the genetic evolution of the coronavirus affects the host response to new variants.
Keywords: COVID-19; FGF23; SpO2/FiO2; TNFRSF11B; machine learning; mortality; ribitol; severity.
Copyright © 2025 Hou, Haslund-Gourley, Diray-Arce, Hoch, Rouphael, Becker, Augustine, Ozonoff, Guan, Kleinstein, Peters, Reed, Altman, Langelier, Maecker, Kim, Montgomery, Krammer, Wilson, Eckalbar, Bosinger, Levy, Steen, Rosen, Baden, Melamed, Ehrlich, McComsey, Sekaly, Schaenman, Shaw, Hafler, Corry, Kheradmand, Atkinson, Brakenridge, Agudelo Higuita, Metcalf, Hough, Messer, Pulendran, Nadeau, Davis, Fernandez Sesma, Simon, Kraft, Bime, Calfee, Erle, IMPACC Network, Robinson, Cairns, Haddad and Comunale.
Conflict of interest statement
The Icahn School of Medicine at Mount Sinai has filed patent applications relating to SARS-CoV-2 serological assays, NDV-based SARS-CoV-2 vaccines, influenza virus vaccines, and influenza virus therapeutics, which list Florian Krammer as co-inventor. Mount Sinai has spun out a company, Kantaro, to market serological tests for SARS-CoV-2 and another company, Castlevax, to develop SARS-CoV-2 vaccines. Florian Krammer is a co-founder and scientific advisory board member of Castlevax. Florian Krammer has consulted for Merck, Curevac, Seqirus, GSK, and Pfizer and is currently consulting for 3rd Rock Ventures, Sanofi, Gritstone, and Avimex. The Krammer laboratory is also collaborating with Dynavax on influenza vaccine development and with VIR on influenza virus therapeutics development. Viviana Simon is a co-inventor on a patent filed relating to SARS-CoV-2 serological assays (the “Serology Assays”). Ofer Levy is a named inventor on patents held by Boston Children's Hospital relating to vaccine adjuvants and human in vitro platforms that model vaccine action. His laboratory has received research support from GlaxoSmithKline (GSK) and is a co-founder of and advisor to Ovax, Inc. Charles Cairns serves as a consultant to bioMerieux and is funded by a grant from the Bill & Melinda Gates Foundation. James A. Overton is a consultant at Knocean Inc. Jessica Lasky-Su serves as a scientific advisor of Precion Inc. Scott R. Hutton, Greg Michelloti, and Kari Wong are employees of Metabolon Inc. Vicki Seyfer-Margolis is a current employee of MyOwnMed. Nadine Rouphael reports grants or contracts with Merck, Sanofi, Pfizer, Vaccine Company, and Immorna, and has participated in data safety monitoring boards and selected advisory boards for Moderna, Sanofi, Seqirus, Pfizer, EMMES, ICON, BARDA, and CyanVan, Imunon Micron. N.R. has also received support for meetings/travel from Sanofi and Moderna and honoraria from Virology Education and Krog Consulting. Adeeb Rahman is a current employee of Immunai Inc. Steven Kleinstein is a consultant related to the ImmPort data repository for Peraton. Nathan Grabaugh is a consultant for Tempus Labs and the National Basketball Association. Akiko Iwasaki is a consultant for 4BIO, Blue Willow Biologics, Revelar Biotherapeutics, RIGImmune, Xanadu Bio, Paratus Sciences. Monika Kraft receives research funds paid to her institution from NIH, ALA, Sanofi, and Astra-Zeneca for work in asthma, serves as a consultant for Astra-Zeneca, Sanofi, Chiesi, and GSK for severe asthma; is a co-founder and CMO for RaeSedo, Inc., a company created to develop peptidomimetics for the treatment of inflammatory lung disease. Esther Melamed received research funding from Babson Diagnostics and an honorarium from the Multiple Sclerosis Association of America and has served on the advisory boards of Genentech, Horizon, Teva, and Viela Bio. Carolyn Calfee receives research funding from NIH, FDA, DOD, Roche-Genentech, and Quantum Leap Healthcare Collaborative, as well as consulting services for Janssen, Vasomune, Gen1e Life Sciences, NGMBio, and Cellenkos. Wade Schulz was an investigator for a research agreement, through Yale University, from the Shenzhen Center for Health Information for work to advance intelligent disease prevention and health promotion; collaborates with the National Center for Cardiovascular Diseases in Beijing; is a technical consultant to Hugo Health, a personal health information platform; cofounder of Refactor Health, an AI-augmented data management platform for healthcare; and has received grants from Merck and Regeneron Pharmaceutical for research related to COVID-19. Grace A McComsey received research grants from Rehdhill, Cognivue, Pfizer, and Genentech, and served as a research consultant for Gilead, Merck, Viiv/GSK, and Janssen. Linda N. Geng received research funding paid to her institution from Pfizer, Inc. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The author(s) declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
Figures


References
-
- World Health Organization . Number of COVID-19 Deaths Reported to WHO (Cumulative Total) (2024). Available online at: https://data.who.int/dashboards/covid19/deaths (Accessed November 31, 2024).
-
- World Health Organization . COVID-19 Vaccine Coverage. World. (2023). Available online at: https://data.who.int/dashboards/covid19/vaccines (Accessed December 31, 2023).
LinkOut - more resources
Full Text Sources
Miscellaneous

