Complex-mediated evasion: modeling defense against antimicrobial peptides with application to human-pathogenic fungus Candida albicans
- PMID: 40695781
- PMCID: PMC12284208
- DOI: 10.1038/s41540-025-00559-1
Complex-mediated evasion: modeling defense against antimicrobial peptides with application to human-pathogenic fungus Candida albicans
Abstract
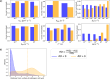
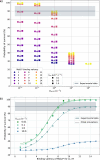
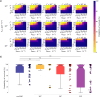
Understanding the complex interplay between host and pathogen during infection is critical for developing diagnostics and improving therapeutic interventions. Among the diverse arsenal employed by the host, antimicrobial peptides (AMP) play a key role in the defense against pathogens. We propose an immune evasion mechanism termed "Complex-mediated evasion" (CME), that allows pathogens to protect themselves against AMP and investigate it through mathematical modeling and computer simulations. To achieve CME, we hypothesize that the pathogen secretes defense molecules that bind AMP. When bound within the complex, AMP are unable to harm the pathogen. Due to molecular gradients, complexes may diffuse away from the pathogen, enhancing the protective effect of the mechanism by decreasing the concentration of AMP in the vicinity of the pathogen. We establish a mathematical model to (i) explore the sensitivity of the mechanism to various parameters and (ii) simulate the immune evasion of the human-pathogenic fungus Candida albicans.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures




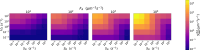


Similar articles
-
Histone deacetylase Sir2 promotes the systemic Candida albicans infection by facilitating its immune escape via remodeling the cell wall and maintaining the metabolic activity.mBio. 2024 Jun 12;15(6):e0044524. doi: 10.1128/mbio.00445-24. Epub 2024 Apr 29. mBio. 2024. PMID: 38682948 Free PMC article.
-
Protein kinase A signaling regulates immune evasion by shaving and concealing fungal β-1,3-glucan.Proc Natl Acad Sci U S A. 2025 Jun 17;122(24):e2423864122. doi: 10.1073/pnas.2423864122. Epub 2025 Jun 9. Proc Natl Acad Sci U S A. 2025. PMID: 40489619 Free PMC article.
-
AI-Driven Antimicrobial Peptide Discovery: Mining and Generation.Acc Chem Res. 2025 Jun 17;58(12):1831-1846. doi: 10.1021/acs.accounts.0c00594. Epub 2025 Jun 3. Acc Chem Res. 2025. PMID: 40459283 Free PMC article. Review.
-
Commensal colonization of Candida albicans in the mouse gastrointestinal tract is mediated via expression of candidalysin and adhesins.Microbiol Spectr. 2025 Jul 30:e0056725. doi: 10.1128/spectrum.00567-25. Online ahead of print. Microbiol Spectr. 2025. PMID: 40736264
-
Anti-Candida albicans natural products, sources of new antifungal drugs: A review.J Mycol Med. 2017 Mar;27(1):1-19. doi: 10.1016/j.mycmed.2016.10.002. Epub 2016 Nov 11. J Mycol Med. 2017. PMID: 27842800
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

