CmFUL1 was potentially involved in fruit elongation in melon
- PMID: 40698194
- PMCID: PMC12282119
- DOI: 10.1093/hr/uhaf138
CmFUL1 was potentially involved in fruit elongation in melon
Abstract
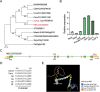
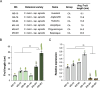
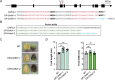
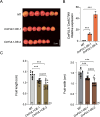
Melon (Cucumis melo L.) is a fruit crop in the world; fruit size and fruit shape are major traits for melon quality. Fruit length is a crucial indicator affecting fruit size and shape, but few genes regulating this trait have been identified. Here, we identified the transcription factor CmFUL1 (FRUITFULL) as a candidate for regulating fruit length using genome-wide association analysis (GWAS) and phylogenetic analysis. CmFUL1 is mainly expressed during flower and ovary development by tissue-specific expression. Transcriptional analysis revealed that CmFUL1 expression levels exhibited a negative correlation with fruit length across diverse melon germplasm. Furthermore, functional characterization demonstrated that CmFUL1 acts as a negative regulator of fruit elongation, CR-Cmful1 mutants generated by CRISPR-Cas9 showing enhanced longitudinal fruits. This repressive role was evolutionarily conserved, as heterologous overexpression of CmFUL1 in tomato consistently inhibited fruit elongation. Collectively, the results suggested that CmFUL1 is a candidate gene involved in regulating fruit length in melon, and provided genetic resources for molecular breeding of melon.
© The Author(s) 2025. Published by Oxford University Press on behalf of Nanjing Agricultural University.
Conflict of interest statement
All authors declare that they have no conflict of interest.
Figures


Similar articles
-
A 13-bp insertion in CmAPRR2 gene disrupts its function in regulating the green rind formation of immature melon fruit (Cucumis melo L.).Plant Sci. 2025 Oct;359:112590. doi: 10.1016/j.plantsci.2025.112590. Epub 2025 Jun 3. Plant Sci. 2025. PMID: 40473118
-
OVATE family gene CmOFP6-19b negatively regulates fruit size in melon (Cucumis melo L.).Hortic Res. 2025 Jun 18;12(9):uhaf148. doi: 10.1093/hr/uhaf148. eCollection 2025 Sep. Hortic Res. 2025. PMID: 40861040 Free PMC article.
-
Validation of CRISPR construct activity and gene function in melon via a hairy root transformation system.Physiol Mol Biol Plants. 2025 May;31(5):753-766. doi: 10.1007/s12298-025-01607-0. Epub 2025 Jun 9. Physiol Mol Biol Plants. 2025. PMID: 40568484
-
Antidepressants for pain management in adults with chronic pain: a network meta-analysis.Health Technol Assess. 2024 Oct;28(62):1-155. doi: 10.3310/MKRT2948. Health Technol Assess. 2024. PMID: 39367772 Free PMC article.
-
The Black Book of Psychotropic Dosing and Monitoring.Psychopharmacol Bull. 2024 Jul 8;54(3):8-59. Psychopharmacol Bull. 2024. PMID: 38993656 Free PMC article. Review.
References
-
- Jeffrey C. A review of the Cucurbitaceae. Bot J Linn Soc. 1980;81:233–47
-
- Pitrat M. Melon genetic resources: phenotypic diversity and horticultural taxonomy. In: Grumet R, Katzir N, Garcia-Mas J, eds. Genetics and Genomics of Cucurbitaceae. Springer International Publishing: Cham, 2017,25–60
-
- Cong B, Barrero LS, Tanksley SD. Regulatory change in YABBY-like transcription factor led to evolution of extreme fruit size during tomato domestication. Nat Genet. 2008;40:800–4 - PubMed
-
- Jenks MA, Bebeli PJ. Breeding for Fruit Quality. Wiley; 2011, 261–279
-
- Monforte AJ, Oliver M, Gonzalo MJ. et al. Identification of quantitative trait loci involved in fruit quality traits in melon (Cucumis melo L.). Theor Appl Genet. 2004;108:750–8 - PubMed
LinkOut - more resources
Full Text Sources

