SMC-like Wadjet system prevents plasmid transfer into Clostridium cellulovorans
- PMID: 40699345
- PMCID: PMC12287153
- DOI: 10.1007/s00253-025-13551-w
SMC-like Wadjet system prevents plasmid transfer into Clostridium cellulovorans
Abstract
This study demonstrates the impact of a Structure Maintenance of Chromosome (SMC)-like Wadjet system on the horizontal gene transfer of plasmids by conjugation to a recipient that naturally containing such a system for the first time. A Clostridium cellulovorans mutant with dramatically improved efficiency to receive plasmid DNA by conjugation was isolated and sequenced. Three spontaneous chromosomal deletions included a type II restriction-modification system, a putative CRISPR system, and a cluster of ORFs named jetABCD encoding a putative Wadjet system. Since nearly nothing is known about the role of naturally occurring Wadjet systems in their native host bacteria, markerless chromosomal deletion of jetABCD in the C. cellulovorans wildtype strain 743B was achieved and the effect on conjugative plasmid uptake was studied. The transconjugation frequency of the jetABCD mutant was increased by about five orders of magnitude compared to wildtype C. cellulovorans recipient cells. Bioinformatic analysis of genome sequences of the Bacillota phylum revealed near-complete mutually exclusive possession of either plasmids < 40 kb or jetABCD genes, indicating high efficiency of Wadjet systems in small plasmid prevention in bacteria. Importantly, the implications of this study go beyond the case of C. cellulovorans. Our study demonstrates that the eradication of Wadjet systems can dramatically improve the uptake of recombinant plasmids and thereby enhance genetic engineering of bacterial strains of interest for biotechnological applications. KEY POINTS: • Native Wadjet system inhibits plasmid transfer by conjugation in C. cellulovorans • Deleting jetABCD increased plasmid uptake by about five orders of magnitude • Possession of Wadjet systems efficiently block plasmid maintenance in Bacillota.
Keywords: Clostridium cellulovorans; CRISPR-Cas; Plasmid defense system; Restriction-modification system; Transconjugation; Wadjet system.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval: This article does not contain any studies with human participants or animals performed by any of the authors. Competing interests: The authors declare no competing interests.
Figures





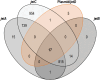
References
-
- Almeida L, Schöllkopf A, Edelmann H, Ehrenreich A, Liebl W (2025) Markerless deletion of the putative type I and III restriction-modification systems in the cellulolytic bacterium Clostridium cellulovorans using a codBA-based counterselection technique. J Biotechnol :22–31. 10.1016/j.jbiotec.2024.11.001 - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

