Dual-Faced Role of GDF6 in Cancer: Mechanistic Insights into Its Context-Dependent Regulation of Metastasis and Immune Evasion Across Human Malignancies
- PMID: 40699648
- PMCID: PMC12025365
- DOI: 10.3390/cimb47040249
Dual-Faced Role of GDF6 in Cancer: Mechanistic Insights into Its Context-Dependent Regulation of Metastasis and Immune Evasion Across Human Malignancies
Abstract
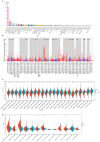
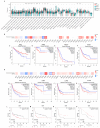
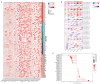
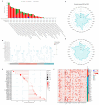
Growth differentiation factor 6 (GDF6), a member of the TGF-β superfamily, plays multifaceted roles in tumorigenesis, yet its molecular mechanisms and cancer-type-specific regulatory networks remain poorly defined. This study investigates GDF6's context-dependent functions through pan-cancer multi-omics integration and functional validation. Transcriptomic data from TCGA (33 cancers, n = 10,535) and GTEx were analyzed to assess GDF6 dysregulation. Co-expression networks, pathway enrichment (KEGG/GO), and epigenetic interactions (m6A, m5C, m1A) were explored. Functional assays included siRNA knockdown, wound healing, and validation in immunotherapy cohorts. GDF6 exhibited bidirectional expression patterns, with downregulation in 23 cancers (e.g., GBM, BRCA) and upregulation in 7 malignancies (e.g., KIRC, PAAD). Mechanistically, GDF6 activated the PI3K-Akt/VEGF pathways, thereby promoting angiogenesis and metastasis. It modulated epigenetic regulation through interactions with m6A readers and erasers. Additionally, GDF6 reshaped the immune microenvironment by recruiting myeloid-derived suppressor cells (MDSCs) and cancer-associated fibroblasts. Notably, GDF6's dual role extended to immunotherapy: it suppressed anti-PD1 efficacy but enhanced anti-PD-L1 sensitivity, linked to differential MHC-II and hypoxia-response regulation. This study deciphers GDF6's context-dependent molecular networks, revealing its dual roles in metastasis and immune evasion. These findings highlight GDF6 as a central node in TGF-β-mediated oncogenic signaling and a potential therapeutic target for precision intervention.
Keywords: GDF6; PI3K-Akt pathway; TGF-β superfamily; epigenetic regulation; immune microenvironment.
Conflict of interest statement
The authors declare no conflicts of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Figures


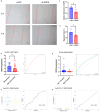
Similar articles
-
Comprehensive pan-cancer analysis reveals NTN1 as an immune infiltrate risk factor and its potential prognostic value in SKCM.Sci Rep. 2025 Jan 25;15(1):3223. doi: 10.1038/s41598-025-85444-x. Sci Rep. 2025. PMID: 39863609 Free PMC article.
-
Integrated pan-cancer analysis of ADM's role in prognosis, immune modulation and resistance.Front Immunol. 2025 Jun 3;16:1573250. doi: 10.3389/fimmu.2025.1573250. eCollection 2025. Front Immunol. 2025. PMID: 40529377 Free PMC article.
-
USP6NL knockdown suppresses colorectal cancer progression by inducing CASP9-Mediated apoptosis and disrupting FOXC2/SNAI1-Driven EMT and angiogenesis.Funct Integr Genomics. 2025 Jul 11;25(1):153. doi: 10.1007/s10142-025-01663-5. Funct Integr Genomics. 2025. PMID: 40643716
-
Hippo/YAP signaling pathway in colorectal cancer: regulatory mechanisms and potential drug exploration.Front Oncol. 2025 Jun 19;15:1545952. doi: 10.3389/fonc.2025.1545952. eCollection 2025. Front Oncol. 2025. PMID: 40612350 Free PMC article. Review.
-
Immune factors and their role in tumor aggressiveness in glioblastoma: Atypical cadherin FAT1 as a promising target for combating immune evasion.Cell Mol Biol Lett. 2025 Jul 25;30(1):89. doi: 10.1186/s11658-025-00769-9. Cell Mol Biol Lett. 2025. PMID: 40713501 Free PMC article. Review.
References
-
- Venkatesan A.M., Vyas R., Gramann A.K., Dresser K., Gujja S., Bhatnagar S., Chhangawala S., Gomes C.B.F., Xi H.S., Lian C.G., et al. Ligand-activated BMP signaling inhibits cell differentiation and death to promote melanoma. J. Clin. Investig. 2018;128:294–308. doi: 10.1172/JCI92513. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

