Protocol to distinguish pre-mRNA from mRNA in RNA-protein interaction studies
- PMID: 40700006
- PMCID: PMC12305230
- DOI: 10.1016/j.xpro.2025.103967
Protocol to distinguish pre-mRNA from mRNA in RNA-protein interaction studies
Abstract
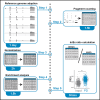
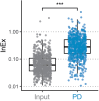
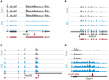
Transcriptome-wide studies on interactions between RNA-binding proteins (RBPs) and protein-coding RNAs in general preclude interpretations regarding RBP preference for binding to the more abundant mRNA over the less abundant pre-mRNA. Here, we present a protocol to determine the binding preference of the RBP tristetraprolin (TTP, Zfp36) for pre-mRNA versus mRNA. We describe steps for the identification and quantitation of intronic and exonic fragments in RNA bound to TTP. This protocol can potentially be applied to any RBP. For complete details on the use and execution of this protocol, please refer to Bestehorn et al.1.
Keywords: Bioinformatics; Gene Expression; Genomics; Molecular Biology; RNA-seq; Sequence analysis; Sequencing.
Copyright © 2025 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures

References
-
- Bestehorn A., von Wirén J., Zeiler C., Fesselet J., Didusch S., Forte M., Doppelmayer K., Borroni M., Le Heron A., Scinicariello S., et al. Cytoplasmic mRNA decay controlling inflammatory gene expression is determined by pre-mRNA fate decision. Mol. Cell. 2025;85:742–755. doi: 10.1016/j.molcel.2025.01.001. - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

