Macrophage-T cell interactions promote SLAMF1 expression for enhanced TB defense
- PMID: 40701970
- PMCID: PMC12287521
- DOI: 10.1038/s41467-025-61826-7
Macrophage-T cell interactions promote SLAMF1 expression for enhanced TB defense
Abstract
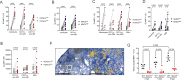
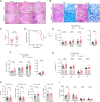
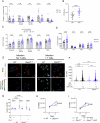
CD4+ T cells are crucial for protective immunity to intracellular pathogens. In addition to secreting cytokines, CD4+ T cells promote control of Mycobacterium tuberculosis infection through cognate interactions with macrophages, but the mechanism has been unclear. Here, we show that SLAMF1/CD150 is highly and uniquely induced in macrophages by antigen-specific interactions with CD4+ T cells. In macrophages, SLAMF1 enhances the generation of reactive oxygen species and restricts Mtb replication. Mtb-infection of mice promotes SLAMF1 expression specifically on infected macrophages, not uninfected bystanders. SLAMF1 expression depends on adaptive immunity and also autophagy. Moreover, Slamf1-/- mice have higher Mtb burden and more rapid disease progression than wild type mice. Using Slamf1fl/fl conditional knock-out mice, we show that in vivo Slamf1 is specifically required in macrophages to restrict mycobacterial growth and limit IL-1β production. In macaques, macrophage SLAMFI expression also correlates with T cell responses and protection. Combined, these data demonstrate that SLAMF1 is a marker of macrophage-T cells interactions, and it promotes protection against Mtb.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures



References
-
- WHO. Global tuberculosis report 2021. (World Health Organization, 2021).
-
- Wolf, A. J. et al. Mycobacterium tuberculosis infects dendritic cells with high frequency and impairs their function in vivo. J. Immunol.179, 2509–2519 (2007). - PubMed
-
- Caruso, A. M. et al. Mice deficient in CD4 T cells have only transiently diminished levels of IFN-gamma, yet succumb to tuberculosis. J. Immunol.162, 5407–5416 (1999). - PubMed
MeSH terms
Substances
Grants and funding
- AI051242/U.S. Department of Health & Human Services | NIH | National Institute of Allergy and Infectious Diseases (NIAID)
- P30 CA016087/CA/NCI NIH HHS/United States
- R01 AI130454/AI/NIAID NIH HHS/United States
- R01 AI184568/AI/NIAID NIH HHS/United States
- AI135098/U.S. Department of Health & Human Services | NIH | National Institute of Allergy and Infectious Diseases (NIAID)
- R21 AI155380/AI/NIAID NIH HHS/United States
- AI184568/U.S. Department of Health & Human Services | NIH | National Institute of Allergy and Infectious Diseases (NIAID)
- AI152321/U.S. Department of Health & Human Services | NIH | National Institute of Allergy and Infectious Diseases (NIAID)
- K08 AI119150/AI/NIAID NIH HHS/United States
- K08 AI135098/AI/NIAID NIH HHS/United States
- AI119150/U.S. Department of Health & Human Services | NIH | National Institute of Allergy and Infectious Diseases (NIAID)
- AI130454/U.S. Department of Health & Human Services | NIH | National Institute of Allergy and Infectious Diseases (NIAID)
- R01 AI051242/AI/NIAID NIH HHS/United States
- F31 AI152321/AI/NIAID NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Research Materials

