De novo mutations mediate phenotypic switching in an opportunistic human lung pathogen
- PMID: 40701980
- PMCID: PMC12287342
- DOI: 10.1038/s41467-025-61168-4
De novo mutations mediate phenotypic switching in an opportunistic human lung pathogen
Abstract
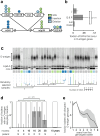
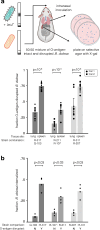
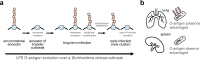
Bacteria evolving within human hosts encounter selective tradeoffs that render mutations adaptive in one context and deleterious in another. Here, we report that the cystic fibrosis-associated pathogen Burkholderia dolosa overcomes in-human selective tradeoffs by acquiring successive point mutations that alternate phenotypes. We sequenced the whole genomes of 931 respiratory isolates from two recently infected cystic fibrosis patients and an epidemiologically-linked, chronically-infected patient. These isolates are contextualized using 112 historical genomes from the same outbreak strain. Within both newly infected patients, convergent mutations that disrupt O-antigen expression quickly arose, comprising 29% and 63% of their B. dolosa communities by 3 years. The selection for loss of O-antigen starkly contrasts with our previous observation of parallel O-antigen-restoring mutations after many years of chronic infection in the historical outbreak. Experimental characterization reveals that O-antigen loss increases uptake in immune cells while decreasing competitiveness in the mouse lung. We propose that the balance of these pressures, and thus whether O-antigen expression is advantageous, depends on tissue localization and infection duration. These results suggest that mutation-driven phenotypic alternation may be underestimated without dense temporal sampling, particularly for microbes with prolonged infection or colonization.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: P.C.B. is a consultant to or holds equity in 10× Genomics, General Automation Lab Technologies/Isolation Bio, Celsius Therapeutics, Next Gen Diagnostics, Cache DNA, Concerto Biosciences, Stately, Ramona Optics, Bifrost Biosystems, and Amber Bio. His laboratory has received research funding from Calico Life Sciences, Merck, and Genentech for unrelated work. The remaining authors declare no competing interests.
Figures


Update of
-
De novo mutations mediate phenotypic switching in an opportunistic human lung pathogen.bioRxiv [Preprint]. 2024 Feb 6:2024.02.06.579193. doi: 10.1101/2024.02.06.579193. bioRxiv. 2024. Update in: Nat Commun. 2025 Jul 23;16(1):6799. doi: 10.1038/s41467-025-61168-4. PMID: 38370793 Free PMC article. Updated. Preprint.
References
-
- Ferenci, T. Trade-off mechanisms shaping the diversity of bacteria. Trends Microbiol.24, 209–223 (2016). - PubMed
-
- Cooper, V. S. & Lenski, R. E. The population genetics of ecological specialization in evolving Escherichia coli populations. Nature407, 736–739 (2000). - PubMed
-
- Juhas, M. Horizontal gene transfer in human pathogens. Crit. Rev. Microbiol.41, 101–108 (2015). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

