The colorectal liver metastasis growth pattern phenotype is not dependent on genotype
- PMID: 40702107
- PMCID: PMC12480539
- DOI: 10.1038/s41416-025-03103-4
The colorectal liver metastasis growth pattern phenotype is not dependent on genotype
Abstract
Background: The histopathological growth patterns (HGPs) of colorectal cancer liver metastases broadly classify patients into two groups post-liver metastasectomy, with encapsulated HGP indicating a more favourable prognosis. The potential association between HGPs and specific mutations is poorly understood.
Methods: Using next-generation sequencing data of 461 resected patients (104 patients with encapsulated versus 357 patients with non-encapsulated HGP), 19 putative colorectal cancer driver genes, tumour mutational burden (TMB), and microsatellite instability (MSI) or POLE mediated hypermutation were compared.
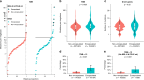
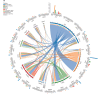
Results: Most putative drivers, including KRAS (q = 0.89), NRAS (q = 0.98),) and BRAF (q = 0.97)), were not associated with HGP. However, mutations in B2M and PTEN were associated with a encapsulated phenotype (7% vs. 0%, q = 0.001, and 9% vs. 2%, q = 0.02, respectively). TMB was higher in encapsulated patients (median 5.8 vs. 5.1 mutations per megabase, p = 0.009). Multivariable overall survival analysis corrected for genetic and patient factors confirmed that the encapsulated phenotype was an independent prognostic factor (adjusted hazard ratio, 0.60; 95% confidence interval: 0.36-0.99). Upon stratified analysis, all identified genetic associations were equivocal between the cohorts.
Conclusions: While an association between genetic drivers of adaptive immune responses seems probable and could explain a minority of encapsulated patients, these results primarily demonstrate that HGP phenotype is independent of the tumour genotype.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests. Ethics approval and consent to participate: All methods were performed in accordance with the relevant guidelines and regulations. Institutional ethical review was obtained from the medical ethics committee of the Erasmus University Medical Center, Rotterdam, the Netherlands (MEC-2018-1743). MSKCC: All the patients signed a clinical (IRB #16-1343 or #15-044) or research (IRB #12-245) consent form for genomic sequencing. New EPOC: The study was approved by the South West Research Ethics Committee, and data were reviewed by the data monitoring and ethics committee. All patients provided written informed consent for further research on their samples during trial enrolment.
Figures



References
-
- Siegel RL, Wagle NS, Cercek A, Smith RA, Jemal A. Colorectal cancer statistics, 2023. CA Cancer J Clin. 2023;73:233–54. - PubMed
-
- Rayner E, van Gool IC, Palles C, Kearsey SE, Bosse T, Tomlinson I, et al. A panoply of errors: polymerase proofreading domain mutations in cancer. Nat Rev Cancer. 2016;16:71–81. - PubMed
-
- Chen DS, Mellman I. Elements of cancer immunity and the cancer–immune set point. Nature. 2017;541:321–30. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

