LncRNA TMEM99 Complexes with IGF2BP2 to Inhibit Autophagy in Lung Adenocarcinoma
- PMID: 40704413
- PMCID: PMC12412623
- DOI: 10.1002/advs.202507871
LncRNA TMEM99 Complexes with IGF2BP2 to Inhibit Autophagy in Lung Adenocarcinoma
Erratum in
-
Correction to "LncRNA TMEM99 Complexes with IGF2BP2 to Inhibit Autophagy in Lung Adenocarcinoma".Adv Sci (Weinh). 2026 Jan;13(2):e21888. doi: 10.1002/advs.202521888. Epub 2025 Nov 27. Adv Sci (Weinh). 2026. PMID: 41309498 Free PMC article. No abstract available.
Abstract
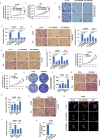
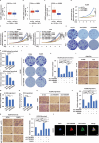
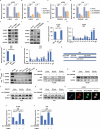
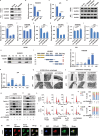
Lung adenocarcinoma (LUAD), the most common type of lung cancer, has a poor prognosis. Long noncoding RNAs (lncRNAs) play a key role in LUAD progression, yet the biological role of lncRNA TMEM99 remains unexplored. In this study, its function in inhibiting autophagy in LUAD is explored. Using RNA sequencing and quantitative reverse transcription PCR (qRT-PCR), TMEM99 is found upregulated in LUAD tissues and cell lines, correlating with poor patient outcomes. In vivo and in vitro assays confirmed that TMEM99 promotes cell proliferation, migration, and invasion, and inhibits autophagy. Mechanistically, the 3' end of TMEM99 binds to the K‑homology domain 1 (KH1) and KH4 domains of far upstream element‑binding protein 3 (FUBP3), stabilizing its protein. The TMEM99-FUBP3 complex binds to p21 mRNA and recruits IGF2BP2 in an N⁶‑methyladenosine (m⁶A)-dependent manner, which enhances mRNA stability and translation efficiency. This study reveals that TMEM99 plays a crucial regulatory role in LUAD autophagy and presents a novel cytoplasmic regulatory mechanism contributing to LUAD progression.
Keywords: FUBP3; Insulin‐like growth factor 2 mRNA‐binding protein 2 (IGF2BP2); TMEM99; autophagy; lung adenocarcinoma.
© 2025 The Author(s). Advanced Science published by Wiley‐VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




References
-
- Bray F., Laversanne M., Sung H., Ferlay J., Siegel R. L., Soerjomataram I., Jemal A., CA Cancer J. Clin. 2024, 74, 229. - PubMed
-
- Zhang Y., Vaccarella S., Morgan E., Li M., Etxeberria J., Chokunonga E., Manraj S. S., Kamate B., Omonisi A., Bray F., Lancet Oncol. 2023, 24, 1206. - PubMed
-
- Makarem M., Janne P. A., Cancer 2024, 130, 3239. - PubMed
-
- Lu S., Wu L., Jian H., Chen Y., Wang Q., Fang J., Wang Z., Hu Y., Sun M., Han L., Miao L., Ding C., Cui J., Li B., Pan Y., Li X., Ye F., Liu A., Wang K., Cang S., Zhou H., Sun X., Ferry D., Lin Y., Wang S., Zhang W., Zhang C., Lancet Oncol. 2022, 23, 1167. - PubMed
MeSH terms
Substances
Grants and funding
- 21Y11913700/Science and Technology Commission of Shanghai Municipality
- 22YF1408900/Shanghai Sailing Program
- Y-2019AZQN-0511/Beijing Xisike Clinical Oncology Research Foundation
- 82172744/National Science Fund for Distinguished Young Scholars
- 82203037/National Science Fund for Distinguished Young Scholars
LinkOut - more resources
Full Text Sources
Medical
