Phage-based delivery of CRISPR-associated transposases for targeted bacterial editing
- PMID: 40711918
- PMCID: PMC12318184
- DOI: 10.1073/pnas.2504853122
Phage-based delivery of CRISPR-associated transposases for targeted bacterial editing
Abstract
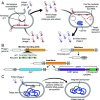
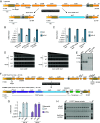
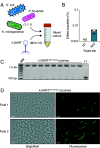
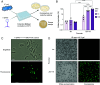
Phage λ, a well-characterized temperate phage, has been recently leveraged for bacterial genome editing by selectively delivering base editors into targeted bacterial species. We extend this concept by engineering phage λ to deliver CRISPR-guided transposases, accomplishing large insertions and targeted gene disruptions. To achieve this, we engineered phage λ using homologous recombination paired with Cas13a-based counterselection for precise phage modifications. Initially, we established the utility of Cas13a in phage λ by conducting minimal recoding edits, deletions, and insertions. Subsequently, we scaled up the engineering to embed the comprehensive DNA-editing CRISPR-Cas transposase (DART) system within the phage genome, creating λ-DART phages. These modified λ-DART phages were then employed to infect Escherichia coli, generating CRISPR RNA-guided transposition events in the host genome. Applying our engineered λ-DART phages to monocultures and a mixed bacterial community comprising three genera led to efficient, precise, and specific gene knockouts and insertions in the targeted E. coli cells, achieving editing efficiencies surpassing 50% of the population. This research enhances phage-mediated genome editing by enabling efficient in situ gene integrations in bacteria, offering an avenue for further application in microbial community contexts. This scalable method enables flexible microbial genome editing in situ to manipulate the function and composition of diverse ecosystems.
Keywords: CRISPR; Cas; genome editing; phage; transposase.
Conflict of interest statement
Competing interests statement:R.B. is a cofounder of Intellia Therapeutics, Locus Biosciences, TreeCo, CRISPR Biotechnologies and Ancilia Biosciences, and a shareholder of Caribou Biosciences, Inari Ag, Tune Therapeutics, and Eligo Biosciences. J.A.D. is a cofounder of Azalea Therapeutics, Caribou Biosciences, Editas Medicine, Scribe Therapeutics, Intellia Therapeutics, and Mammoth Biosciences. J.A.D. is an advisor to Vertex, Scribe Therapeutics, Algen Biotechnologies, The Column Group, Inari, Sixth Street, Johnson & Johnson, Altos and Tempus.
Figures

Similar articles
-
A recombineering-based platform for high-throughput genomic editing in Escherichia coli.Appl Environ Microbiol. 2025 Jul 23;91(7):e0019325. doi: 10.1128/aem.00193-25. Epub 2025 Jun 12. Appl Environ Microbiol. 2025. PMID: 40503884 Free PMC article.
-
Bacterial genome engineering using CRISPR-associated transposases.Nat Protoc. 2024 Mar;19(3):752-790. doi: 10.1038/s41596-023-00927-3. Epub 2024 Jan 12. Nat Protoc. 2024. PMID: 38216671 Free PMC article. Review.
-
Characterization of targeted knock-in achieved via tandem paired nicking mediated by CRISPR/Cas9 nickases.Methods. 2025 Sep;241:184-195. doi: 10.1016/j.ymeth.2025.06.004. Epub 2025 Jun 10. Methods. 2025. PMID: 40505881
-
Continuous multiplexed phage genome editing using recombitrons.Nat Biotechnol. 2025 Aug;43(8):1299-1310. doi: 10.1038/s41587-024-02370-5. Epub 2024 Sep 5. Nat Biotechnol. 2025. PMID: 39237706 Free PMC article.
-
Genome-Editing Tools for Lactic Acid Bacteria: Past Achievements, Current Platforms, and Future Directions.Int J Mol Sci. 2025 Aug 2;26(15):7483. doi: 10.3390/ijms26157483. Int J Mol Sci. 2025. PMID: 40806610 Free PMC article. Review.
References
-
- Bagarolli R. A., et al. , Probiotics modulate gut microbiota and improve insulin sensitivity in DIO mice. J. Nutr. Biochem. 50, 16–25 (2017). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

