Recognising, quantifying and accounting for classification uncertainty in type 2 diabetes subtypes
- PMID: 40715719
- PMCID: PMC12423224
- DOI: 10.1007/s00125-025-06486-4
Recognising, quantifying and accounting for classification uncertainty in type 2 diabetes subtypes
Abstract
Aims/hypothesis: Despite continued interest in precision diagnostics and type 2 diabetes subtypes, the challenge of uncertainty in the classification of individuals into subtypes remains. This study introduces a novel method for quantifying and accounting for classification uncertainty in type 2 diabetes subtypes.
Methods: Building on recommendations from the ADA/EASD Precision Medicine in Diabetes Initiative, we quantified classification uncertainty using the normalised relative entropy (NRE), computed from distances to cluster centroids. A lower NRE value indicates greater uncertainty in an individual's cluster assignment. We examined the NRE in a cohort of 859 individuals with recent-onset type 2 diabetes from the prospective, observational German Diabetes Study (GDS) and compared it across previously identified diabetes subtypes, defined by age, BMI, HbA1c, HOMA-IR and HOMA-B. Predicted 10 year CVD risk (SCORE2-Diabetes) of the subtypes was evaluated with and without accounting for classification uncertainty.
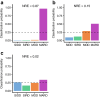
Results: Individuals with mild age-related diabetes (n=395) and mild obesity-related diabetes (n=316) had a median NRE of 0.155 (95% CI 0.142, 0.177) and 0.119 (95% CI 0.107, 0.131), respectively. By contrast, individuals with severe insulin-resistant diabetes (n=130) and severe insulin-deficient diabetes (n=18) had a lower median NRE of 0.086 (95% CI 0.075, 0.108) and 0.082 (95% CI 0.071, 0.109), respectively. After weighting individuals by classification certainty, the proportion of variation in SCORE2-Diabetes explained by the subtypes (R2) increased from 17.4% (95% CI 12.8, 23.0) to 31.5% (95% CI 26.4, 37.1). The predicted 10 year CVD risk of the mild age-related diabetes subtype increased from 10.3% (95% CI 9.8, 10.7) to 11.6% (95% CI 11.2, 12.0).
Conclusions/interpretation: The NRE provides a means to quantify and compare individual classification uncertainty in type 2 diabetes subtypes. Classification uncertainty varied between subtypes and individuals with type 2 diabetes, and accounting for it improved the ability of the subtypes to predict 10 year CVD risk.
Keywords: Classification uncertainty; Clusters; German Diabetes Study; Precision medicine; Relative entropy; Subtypes; Type 2 diabetes mellitus.
© 2025. The Author(s).
Conflict of interest statement
Acknowledgements: The German Diabetes Study (GDS) Group consists of Michael Roden (speaker)1,2,3, Hadi Al-Hasani3,4, Bengt Frederik Belgardt3,5, Gidon J Bönhof1,2,3, Gerd Geerling6, Christian Herder1,2,3, Andrea Icks3,7,8, Karin Jandeleit-Dahm1,9, Jorg Kotzka3,4, Oliver Kuß3,10, Eckhard Lammert3,5,11, Wolfgang Rathmann3,10, Sabrina Schlesinger3,10, Vera Schrauwen-Hinderling1,3,, Julia Szendroedi12, Sandra Trenkamp1,3, Robert Wagner1,2,3 and their co-workers who are responsible for the design and conduct of the GDS. 1 Institute for Clinical Diabetology, German Diabetes Center, Leibniz Center for Diabetes Research at Heinrich Heine University, Düsseldorf, Germany 2 Division of Endocrinology and Diabetology, Medical Faculty and University Hospital Düsseldorf, Heinrich Heine University Düsseldorf, Düsseldorf, Germany 3 German Center for Diabetes Research (DZD), München-Neuherberg, Germany 4 Institute for Clinical Biochemistry and Pathobiochemistry, German Diabetes Center, Leibniz Center for Diabetes Research at Heinrich Heine University Düsseldorf, Düsseldorf, Germany 5 Institute for Vascular and Islet Cell Biology, German Diabetes Center, Leibniz Center for Diabetes Research at Heinrich Heine University Düsseldorf, Düsseldorf, Germany 6 Department of Ophthalmology, Medical Faculty and University Hospital Düsseldorf, Heinrich Heine University Düsseldorf, Düsseldorf, Germany 7 Institute for Health Services Research and Health Economics, German Diabetes Center, Leibniz Center for Diabetes Research at Heinrich Heine University Düsseldorf, Düsseldorf, Germany 8 Institute for Health Services Research and Health Economics, Centre for Health and Society, Medical Faculty and University Hospital Düsseldorf, Heinrich Heine University Düsseldorf, Düsseldorf, Germany 9 Department of Diabetes, Central Clinical School, Monash University, Melbourne, Victoria, Australia 10 Institute for Biometrics and Epidemiology, German Diabetes Center, Leibniz Center for Diabetes Research at Heinrich Heine University Düsseldorf, Düsseldorf, Germany 11 Institute of Metabolic Physiology, Faculty of Mathematics and Natural Sciences, Heinrich Heine University Düsseldorf, Düsseldorf, Germany 12 Department of Medicine I and Clinical Chemistry, University Hospital of Heidelberg, Heidelberg, Germany Data availability: The dataset analysed in this study is not publicly accessible due to national data protection regulations and restrictions set by the ethics committee to safeguard the privacy of study participants. However, access can be requested through a specific project agreement with the principal investigator of the German Diabetes Study (GDS) (michael.roden@ddz.de). The study protocol and detailed methodology are published in the cohort profile [14] and are freely available. Code availability statement: The R code of this study is available upon reasonable request from the corresponding author. Funding: Open Access funding enabled and organized by Projekt DEAL. The German Diabetes Study (GDS) was initiated and financed by the German Diabetes Center (which is funded by the German Federal Ministry of Health and the Ministry of Culture and Science of the state of North Rhine-Westphalia), the German Diabetes Association, the German Federal Ministry of Education and Research (to the German Center for Diabetes Research, DZD) and the Schmutzler Stiftung. JMD is supported by a Wellcome Trust Early-Career Award (227070/Z/23/Z) and the NIHR Exeter Biomedical Research Centre. The views expressed are those of the authors and not necessarily those of the NIHR or the Department of Health and Social Care. The research of MR is further supported by grants from the European Community (HORIZON-HLTH-2022-STAYHLTH-02-01: Panel A) to the INTERCEPT-T2D consortium and the German Science Foundation (DFG; RTG/GRK 2576 vivid, Project 3 and 493659010 Future4CSPMM). NS is supported by the German Center for Diabetes Research, DZD. The research of OPZ is supported by grants from the European Foundation for the Study of Diabetes (EFSD Rising Star Award) and the German Diabetes Association (DDG Adam Heller Prize). The project has received funding from the programme ‘Profilbildung 2020’, an initiative of the Ministry of Culture and Science of the State of North Rhine-Westphalia (PROFILNRW-2020-107-B). The current research of SK is supported by J. Rettenmaier & Söhne (Rosenberg, Germany), the Wilhelm-Doerenkamp-Foundation (Chur, Switzerland), and the Almond Board of California (Modesto, USA). SM is supported by the ‘Advanced Clinician Scientist Program’ from the University of Lübeck. TM is supported by the EASD mentorship programme. The sole responsibility for the content of this publication lies with the authors. The funding sources had no role in study design, data collection, data analysis, data interpretation, or writing of the report. Authors’ relationships and activities: KM received fees for consulting or lecturing from Eli Lilly, Novo Nordisk and Oviva, and performed investigator-initiated research with support from Sanofi-Aventis. MB received honoraria as a consultant and speaker from Amgen, AstraZeneca, Bayer, Boehringer Ingelheim, Daiichi Sankyo, Lilly, MSD, Novo Nordisk and Sanofi. MR received fees for consulting or lecturing from Astra Zeneca, Boehringer Ingelheim, Echosens, Eli Lilly, Madrigal, MSD, Novo Nordisk and Pfizer, and performed investigator-initiated research with support from Boehringer Ingelheim and Novo Nordisk. NS received fees for consulting or lecturing from Allergan, AstraZeneca, Boehringer Ingelheim, Gilead, Genkyotex, GSK, Intercept Pharma, Lilly, Merck Sharpe & Dohme, Novartis, Novo Nordisk, Pfizer and Sanofi, and received research support from AstraZeneca, Sanofi, DSM Nutritional Products and Roche Diagnostics. OPZ reports receiving lecture fees from Sanofi and Chiesi. RW reports receiving lecture fees from Novo Nordisk, Sanofi-Aventis, Boehringer Ingelheim and Eli Lilly, and has served on the advisory board for Akcea Therapeutics, Daiichi Sankyo, Sanofi-Aventis, Eli Lilly and NovoNordisk. SK reports receiving lecture fees from Berlin-Chemie, Lilly, Boehringer Ingelheim and JuZo-Akademie. SM received lecture fees from Eli Lilly and Novo Nordisk, and has participated on advisory boards for Novo Nordisk. SM’s husband is an employee at Novo Nordisk. The authors declare that there are no other relationships or activities that might bias, or be perceived to bias, their work. Contribution statement: TM contributed to the conceptualisation, methodology, software development, formal analysis, and writing of the original draft, as well as reviewing and editing the manuscript. OPZ, KS, JD, KM, SK, SB, JS, MB, SM, JS, AB, NS and MR contributed substantially to the interpretation of data and reviewed the article critically for important intellectual content. RW contributed to the conceptualisation, participated in reviewing and editing the manuscript and provided supervision. OK contributed to the conceptualisation and methodology, participated in reviewing and editing the manuscript, and provided supervision. All authors gave final approval of this version to be published. TM is the guarantor of this work and, as such, had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Figures





References
-
- Ahlqvist E, Storm P, Käräjämäki A et al (2018) Novel subgroups of adult-onset diabetes and their association with outcomes: a data-driven cluster analysis of six variables. Lancet Diabetes Endocrinol 6(5):361–369. 10.1016/s2213-8587(18)30051-2 - PubMed
-
- Zaharia OP, Strassburger K, Strom A et al (2019) Risk of diabetes-associated diseases in subgroups of patients with recent-onset diabetes: a 5-year follow-up study. Lancet Diabetes Endocrinol 7(9):684–694. 10.1016/S2213-8587(19)30187-1 - PubMed
-
- Van Smeden M, Harrell FE, Dahly DL (2018) Novel diabetes subgroups. Lancet Diabetes Endocrinol 6(6):439–440. 10.1016/S2213-8587(18)30124-4 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- Trust Early-Career Award (227070/Z/23/Z)/WT_/Wellcome Trust/United Kingdom
- EFSD Rising Star Award/European Foundation for the Study of Diabetes
- DDG Adam Heller Prize/Deutsche Diabetes Gesellschaft
- HORIZON-HLTH-2022-STAYHLTH-02-01: Panel A/Horizon Europe Cluster 1: Health
- Profilbildung 2020 (PROFILNRW-2020-107-B)/Ministerium für Kultur und Wissenschaft des Landes Nordrhein-Westfalen
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

