Prediction of 5-year postoperative survival and analysis of key prognostic factors in stage III colorectal cancer patients using novel machine learning algorithms
- PMID: 40727467
- PMCID: PMC12301204
- DOI: 10.3389/fonc.2025.1604386
Prediction of 5-year postoperative survival and analysis of key prognostic factors in stage III colorectal cancer patients using novel machine learning algorithms
Abstract
Objective: This study explores the predictive value of clinical and socio-demographic characteristics for postoperative survival in stage III colorectal cancer (CRC) patients and develops a 5-year postoperative survival prediction model using machine learning algorithms.
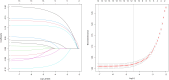
Methods: Data from 13,855 stage III CRC patients who underwent surgery were extracted from the SEER database. Key variables, including marital status, gender, tumor location, histological type, T stage, chemotherapy status, age, tumor size, lymph node ratio, and others, were collected. Data were split into a 7:3 training-validation ratio. Optimal cutoff points for age, tumor diameter, and lymph node ratio were determined using X-tile software. Independent prognostic factors for postoperative survival in stage III colorectal cancer patients were identified through univariate and multivariate logistic regression as well as Lasso regression analyses. These factors were incorporated into machine learning models, including logistic regression, decision tree, LightGBM, and others. ROC curves, calibration curves, and decision curve analysis were used to assess model performance. External validation was performed using data from Shanxi Bethune Hospital.
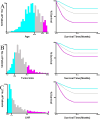
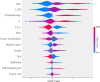
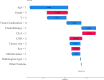
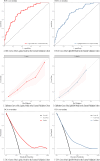
Results: Optimal cutoff points were identified for age (65, 80 years), tumor size (29 mm, 74 mm), and lymph node ratio (0.11, 0.49). Both multivariate logistic regression and Lasso regression consistently identified marital status, tumor location, histological type, T stage, chemotherapy, radiotherapy, age, maximum tumor diameter, lymph node ratio, serum carcinoembryonic antigen (CEA) level, perineural invasion, and tumor differentiation as independent prognostic factors for 5-year postoperative survival in patients with stage III colorectal cancer (P < 0.05). The models showed excellent predictive performance with AUC values ranging from 0.766 to 0.791 in the validation cohort. Age, lymph node ratio, chemotherapy, and T stage were key factors. External validation confirmed model accuracy and clinical applicability.
Conclusion: This study developed and validated an interpretable machine learning model that predicts the 5-year postoperative survival of stage III CRC patients, offering potential for personalized treatment plans.
Keywords: SEER database; colorectal cancer; machine learning; prognostic model; survival prognosis.
Copyright © 2025 Zhang, Li, Jia, Yang, Hu, Wang and Wang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures






Similar articles
-
Competing risk and random survival forest models for predicting survival in post-resection elderly stage I-III colorectal cancer patients.Sci Rep. 2025 Jul 7;15(1):24269. doi: 10.1038/s41598-025-05824-1. Sci Rep. 2025. PMID: 40624131 Free PMC article.
-
Comparison of Two Modern Survival Prediction Tools, SORG-MLA and METSSS, in Patients With Symptomatic Long-bone Metastases Who Underwent Local Treatment With Surgery Followed by Radiotherapy and With Radiotherapy Alone.Clin Orthop Relat Res. 2024 Dec 1;482(12):2193-2208. doi: 10.1097/CORR.0000000000003185. Epub 2024 Jul 23. Clin Orthop Relat Res. 2024. PMID: 39051924
-
Are Current Survival Prediction Tools Useful When Treating Subsequent Skeletal-related Events From Bone Metastases?Clin Orthop Relat Res. 2024 Sep 1;482(9):1710-1721. doi: 10.1097/CORR.0000000000003030. Epub 2024 Mar 22. Clin Orthop Relat Res. 2024. PMID: 38517402
-
Cost-effectiveness of using prognostic information to select women with breast cancer for adjuvant systemic therapy.Health Technol Assess. 2006 Sep;10(34):iii-iv, ix-xi, 1-204. doi: 10.3310/hta10340. Health Technol Assess. 2006. PMID: 16959170
-
Impact of residual disease as a prognostic factor for survival in women with advanced epithelial ovarian cancer after primary surgery.Cochrane Database Syst Rev. 2022 Sep 26;9(9):CD015048. doi: 10.1002/14651858.CD015048.pub2. Cochrane Database Syst Rev. 2022. PMID: 36161421 Free PMC article.
References
-
- Dienstmann R, Mason MJ, Sinicrope FA, Phipps AI, Tejpar S, Nesbakken A, et al. Prediction of overall survival in stage II and III colon cancer beyond TNM system: a retrospective, pooled biomarker study. Ann oncology: Off J Eur Soc Med Oncol. (2017) 28:1023–31. doi: 10.1093/annonc/mdx052, PMID: - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources

