Identification of hub gene associated with colorectal cancer: Integrating Mendelian randomization, transcriptome analysis and experimental verification
- PMID: 40729369
- PMCID: PMC12349882
- DOI: 10.1371/journal.pgen.1011788
Identification of hub gene associated with colorectal cancer: Integrating Mendelian randomization, transcriptome analysis and experimental verification
Abstract
Background: Colorectal cancer (CRC) is a prevalent malignancy with significant mortality rates globally. Understanding the genetic and molecular mechanisms underlying CRC development is crucial for improving therapeutic strategies.
Method: In this study, we utilized cis-eQTL summary data to identify genes potentially causally associated with CRC. The expression levels of candidate genes in tumor and normal tissues were compared using the GEPIA2 database. The correlations between FUT8 expression and cellular functions, tumor mutation burden, immune checkpoint genes, and immune infiltration were analyzed. Molecular docking was performed to identify potential drugs targeting FUT8, and the effects of the selected drug on cell proliferation were evaluated using the MTT assay. Additionally, the cellular thermal shift assay (CETSA) was employed to assess the interaction between the drug and the target protein.
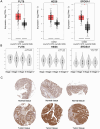
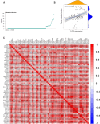
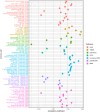
Results: We identified 19 genes with eQTLs potentially associated with CRC, among which six eQTLs were associated with increased CRC risk, including FUT8. FUT8 was significantly overexpressed in CRC tumor tissues and correlated with various cellular functions such as stemness, invasion, EMT, and metastasis. Higher FUT8 expression was associated with higher tumor mutation burden and significant correlations with multiple immune checkpoint genes. Molecular docking identified VE-822 as a promising drug candidate targeting FUT8, which demonstrated inhibitory effects on CRC cell proliferation. The CETSA results indicated that VE ‒ 822 could bind to FUT8 and improve its thermal stability.
Conclusion: FUT8 is a crucial gene that causes colon cancer and is linked to tumour immunity. VE-822 is a promising candidate for treating CRC by targeting FUT8.
Copyright: © 2025 Zhang et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



Similar articles
-
Potential of SPHK1 as a prognostic marker and therapeutic target in colorectal cancer: insights from bioinformatics and experimental analysis.Int J Surg. 2025 Jul 1;111(7):4550-4575. doi: 10.1097/JS9.0000000000002506. Epub 2025 May 28. Int J Surg. 2025. PMID: 40434736
-
Resveratrol suppresses liver cancer progression by downregulating AKR1C3: targeting HCC with HSA nanomaterial as a carrier to enhance therapeutic efficacy.Apoptosis. 2024 Oct;29(9-10):1429-1453. doi: 10.1007/s10495-024-01995-w. Epub 2024 Jul 18. Apoptosis. 2024. PMID: 39023830
-
Comprehensive analysis of Mendelian randomization and scRNA-seq identify key prognostic genes and relevant functional roles in colorectal cancer.Sci Rep. 2025 Jul 11;15(1):25039. doi: 10.1038/s41598-025-10354-x. Sci Rep. 2025. PMID: 40646181 Free PMC article.
-
Systemic treatments for metastatic cutaneous melanoma.Cochrane Database Syst Rev. 2018 Feb 6;2(2):CD011123. doi: 10.1002/14651858.CD011123.pub2. Cochrane Database Syst Rev. 2018. PMID: 29405038 Free PMC article.
-
Guaiac-based faecal occult blood tests versus faecal immunochemical tests for colorectal cancer screening in average-risk individuals.Cochrane Database Syst Rev. 2022 Jun 6;6(6):CD009276. doi: 10.1002/14651858.CD009276.pub2. Cochrane Database Syst Rev. 2022. PMID: 35665911 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

