Integrated Transcriptomic and Proteomic Analyses Reveal CsrA-Mediated Regulation of Virulence and Metabolism in Vibrio alginolyticus
- PMID: 40732024
- PMCID: PMC12298160
- DOI: 10.3390/microorganisms13071516
Integrated Transcriptomic and Proteomic Analyses Reveal CsrA-Mediated Regulation of Virulence and Metabolism in Vibrio alginolyticus
Abstract
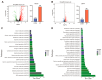
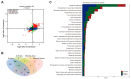
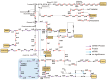
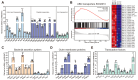
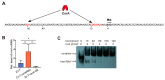
Vibrio alginolyticus, a common Gram-negative opportunistic pathogen of marine animals and humans, is known for its rapid growth in organic-matter-rich environments. However, it remains unclear how it incorporates metabolic pathways in response to diverse carbon and nitrogen sources and rapidly alters gene expression. Increasing evidence suggests that post-transcriptional regulation by RNA-binding proteins and small RNAs (sRNAs) plays a crucial role in bacterial adaptation and metabolism. CsrA (carbon storage regulator A), a conserved post-transcriptional regulator in Gammaproteobacteria, is poorly characterized in Vibrio species. Using integrated transcriptomic and proteomic analyses, we found that CsrA alters the expression of 661 transcripts and 765 protein transcripts in V. alginolyticus, influencing key pathways including central carbon metabolism, amino acid metabolism and transport, quorum sensing, and bacterial secretion systems. Through directed CsrA-RNA EMSAs, we identified several direct mRNA targets of CsrA, including gltB, gcvP, aceE, and tdh, as well as secretion system components (tagH, tssL, yopD, and sctC). Notably, CsrA also directly regulates rraA, a key modulator of ribonuclease activity, suggesting a broader role in RNA metabolism. Our findings establish CsrA as a global regulator in V. alginolyticus, expanding the known targets of CsrA and providing new insights into its regulatory roles.
Keywords: CsrA; Vibrio alginolyticus; amino acid metabolism; central carbon metabolism; transcription factor; virulence.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures


Similar articles
-
Post-Transcriptional Regulation of the MiaA Prenyl Transferase by CsrA and the Small RNA CsrB in Escherichia coli.Int J Mol Sci. 2025 Jun 24;26(13):6068. doi: 10.3390/ijms26136068. Int J Mol Sci. 2025. PMID: 40649849 Free PMC article.
-
CsrA-mediated regulation of a virulence switch in Acinetobacter baumannii.mBio. 2025 Apr 9;16(4):e0405824. doi: 10.1128/mbio.04058-24. Epub 2025 Feb 25. mBio. 2025. PMID: 39998216 Free PMC article.
-
Regulatory mechanism for host-cell contact-dependent T3SS gene expression in Vibrio parahaemolyticus.mSystems. 2025 Jul 22;10(7):e0025125. doi: 10.1128/msystems.00251-25. Epub 2025 Jun 17. mSystems. 2025. PMID: 40525855 Free PMC article.
-
Factors that influence parents' and informal caregivers' views and practices regarding routine childhood vaccination: a qualitative evidence synthesis.Cochrane Database Syst Rev. 2021 Oct 27;10(10):CD013265. doi: 10.1002/14651858.CD013265.pub2. Cochrane Database Syst Rev. 2021. PMID: 34706066 Free PMC article.
-
Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis.Cochrane Database Syst Rev. 2020 Jan 9;1(1):CD011535. doi: 10.1002/14651858.CD011535.pub3. Cochrane Database Syst Rev. 2020. Update in: Cochrane Database Syst Rev. 2021 Apr 19;4:CD011535. doi: 10.1002/14651858.CD011535.pub4. PMID: 31917873 Free PMC article. Updated.
References
Grants and funding
LinkOut - more resources
Full Text Sources

