Population Structure, Genomic Features, and Antibiotic Resistance of Avian Pathogenic Escherichia coli in Shandong Province and Adjacent Regions, China (2008-2023)
- PMID: 40732164
- PMCID: PMC12298790
- DOI: 10.3390/microorganisms13071655
Population Structure, Genomic Features, and Antibiotic Resistance of Avian Pathogenic Escherichia coli in Shandong Province and Adjacent Regions, China (2008-2023)
Abstract
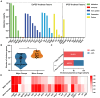
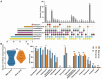
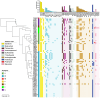
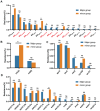
Avian pathogenic Escherichia coli (APEC) poses a global threat to poultry health and public safety due to its high lethality, limited treatment options, and potential for zoonotic transmission via the food chain. However, long-term genomic surveillance remains limited, especially in countries like China where poultry farming is highly intensive. This study aimed to characterize the population structure, virulence traits, and antimicrobial resistance of 81 APEC isolates from diseased chickens collected over 16 years from Shandong and neighboring provinces in eastern China. The isolates were grouped into seven Clermont phylogroups, with A and B1 being dominant. MLST revealed 27 STs, and serotyping identified 29 O and 16 H antigens, showing high genetic diversity. The minor phylogroups (B2, C, D, E, G) encoded more virulence genes and had higher virulence-plasmid ColV carriage, with enrichment for iron-uptake, protectins, and extraintestinal toxins. In contrast, the dominant phylogroups A and B1 primarily carried adhesin and enterotoxin genes. Antimicrobial resistance was widespread: 76.5% of isolates were multidrug-resistant. The minor phylogroups exhibited higher tetracycline resistance (mediated by tet(A)), whereas the major phylogroups showed increased resistance to third- and fourth-generation cephalosporins (due to blaCTX-M-type ESBL genes). These findings offer crucial data for APEC prevention and control, safeguarding the poultry industry and public health.
Keywords: antimicrobial resistance; avian pathogenic Escherichia coli; population structure; virulence; whole-genome sequencing.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures

Similar articles
-
Prevalence of specific serogroups, antibiotic resistance and virulence factors of avian pathogenic Escherichia coli (APEC) isolated from clinical cases: A systematic review and meta-analysis.Microb Pathog. 2024 Sep;194:106843. doi: 10.1016/j.micpath.2024.106843. Epub 2024 Aug 6. Microb Pathog. 2024. PMID: 39117015
-
Antimicrobial resistance and virulence gene profiles of Escherichia coli isolated from poultry farms using One Health perspective in Abeokuta, Nigeria.BMC Microbiol. 2025 Jul 16;25(1):440. doi: 10.1186/s12866-025-04135-9. BMC Microbiol. 2025. PMID: 40670940 Free PMC article.
-
Whole-genome characterisation of Escherichia coli isolates from patients with bacteraemia presenting with sepsis or septic shock in Spain: a multicentre cross-sectional study.Lancet Microbe. 2024 Apr;5(4):e390-e399. doi: 10.1016/S2666-5247(23)00369-5. Epub 2024 Mar 26. Lancet Microbe. 2024. PMID: 38547882
-
Phenotypic and genotypic characterization of biofilm-producing avian pathogenic Escherichia coli (APEC) isolates from Algerian poultry: associations between antimicrobial resistance and virulence genes.Vet Res Commun. 2025 Jun 21;49(4):232. doi: 10.1007/s11259-025-10801-0. Vet Res Commun. 2025. PMID: 40542899
-
Strategic Detection of Escherichia coli in the Poultry Industry: Food Safety Challenges, One Health Approaches, and Advances in Biosensor Technologies.Biosensors (Basel). 2025 Jul 1;15(7):419. doi: 10.3390/bios15070419. Biosensors (Basel). 2025. PMID: 40710069 Free PMC article. Review.
References
-
- Swelum A.A., Elbestawy A.R., El-Saadony M.T., Hussein E.O., Alhotan R., Suliman G.M., Taha A.E., Ba-Awadh H., El-Tarabily K.A., El-Hack M.E.A. Ways to minimize bacterial infections, with special reference to Escherichia coli, to cope with the first-week mortality in chicks: An updated overview. Poult. Sci. 2021;100:101039. doi: 10.1016/j.psj.2021.101039. - DOI - PMC - PubMed
Grants and funding
- 2023YFD1800805/the National Key Research and Development Program of China
- 2024YFE0111700/the National Key Research and Development Program of China
- tsqn202408303/the Taishan Scholars Program
- CXGC2025F10/Agricultural Science and Technology Innovation Project of Shandong Academy of Agricultural Sciences
LinkOut - more resources
Full Text Sources

