Flipping the Target: Evaluating Natural LDHA Inhibitors for Selective LDHB Modulation
- PMID: 40733189
- PMCID: PMC12299506
- DOI: 10.3390/molecules30142923
Flipping the Target: Evaluating Natural LDHA Inhibitors for Selective LDHB Modulation
Abstract
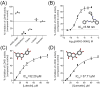
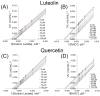
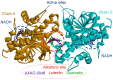
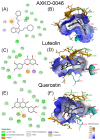
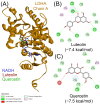
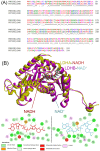
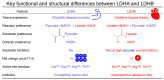
Lactate dehydrogenase (LDH) catalyzes the reversible interconversion of pyruvate and lactate, coupled with the redox cycling of NADH and NAD+. While LDHA has been extensively studied as a therapeutic target, particularly in cancer, due to its role in the Warburg effect, LDHB remains underexplored, despite its involvement in the metabolic reprogramming of specific cancer types, including breast and lung cancers. Most known LDH inhibitors are designed against the LDHA isoform and act competitively at the active site. In contrast, LDHB exhibits distinct kinetic properties, substrate preferences, and structural features, warranting isoform-specific screening strategies. In this study, 115 natural compounds previously reported as LDHA inhibitors were systematically evaluated for LDHB inhibition using an integrated in silico and in vitro approach. Virtual screening identified 16 lead phytochemicals, among which luteolin and quercetin exhibited uncompetitive inhibition of LDHB, as demonstrated by enzyme kinetic assays. These findings were strongly supported by molecular docking analyses, which revealed that both compounds bind at an allosteric site located at the dimer interface, closely resembling the binding mode of the established LDHB uncompetitive inhibitor AXKO-0046. In contrast, comparative docking against LDHA confirmed their active-site binding and competitive inhibition, underscoring their isoform-specific behavior. Our findings highlight the necessity of assay conditions tailored to LDHB's physiological role and demonstrate the application of a previously validated colorimetric assay for high-throughput screening. This work lays the foundation for the rational design of selective LDHB inhibitors from natural product libraries.
Keywords: allosteric inhibition; cancer metabolism; isoform selectivity; lactate dehydrogenase B; natural products.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures

Similar articles
-
Identification of the first highly selective inhibitor of human lactate dehydrogenase B.Sci Rep. 2021 Nov 1;11(1):21353. doi: 10.1038/s41598-021-00820-7. Sci Rep. 2021. PMID: 34725423 Free PMC article.
-
Exploring Type II Diabetes Inhibitors from Genus Daphne Plant-species: An Integrated Computational Study.Comb Chem High Throughput Screen. 2025;28(8):1413-1442. doi: 10.2174/0113862073262227231005074024. Comb Chem High Throughput Screen. 2025. PMID: 38584562
-
The Black Book of Psychotropic Dosing and Monitoring.Psychopharmacol Bull. 2024 Jul 8;54(3):8-59. Psychopharmacol Bull. 2024. PMID: 38993656 Free PMC article. Review.
-
Lactate dehydrogenase A-coupled NAD+ regeneration is critical for acute myeloid leukemia cell survival.Cancer Metab. 2025 May 19;13(1):22. doi: 10.1186/s40170-025-00392-4. Cancer Metab. 2025. PMID: 40390151 Free PMC article.
-
Management of urinary stones by experts in stone disease (ESD 2025).Arch Ital Urol Androl. 2025 Jun 30;97(2):14085. doi: 10.4081/aiua.2025.14085. Epub 2025 Jun 30. Arch Ital Urol Androl. 2025. PMID: 40583613 Review.
References
-
- Farhana A., Lappin S.L. StatPearls. StatPearls Publishing; Treasure Island, FL, USA: 2022. Biochemistry, lactate dehydrogenase. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

