Design and Biological Evaluation of hBest1-Containing Bilayer Nanostructures
- PMID: 40733213
- PMCID: PMC12300095
- DOI: 10.3390/molecules30142948
Design and Biological Evaluation of hBest1-Containing Bilayer Nanostructures
Abstract
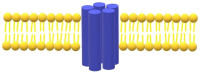
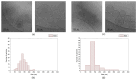
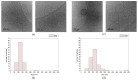
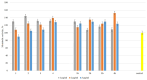
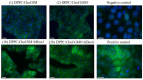
Bestrophinopathies are a group of inherited retinal diseases caused by mutations in the BEST1 gene. The protein encoded by this gene, bestorphin-1 (hBest1), is a calcium-dependent transmembrane channel localized on the basolateral membrane of retinal pigment epithelial (RPE) cells. We have already demonstrated the surface behavior and organization of recombinant hBest1 and its interactions with membrane lipids such as 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (POPC), sphingomyelin (SM) and cholesterol (Chol) in models of biological membranes, which affect the hBest1 structure-function relationship. The main aim of our current investigation is to integrate pure hBest1 protein into lipid bilayer nanostructures. We synthesized and characterized various hBest1-containing nanostructures based on 1,2-Dipalmitoylphosphatidylcholine (DPPC), SM, glycerol monooleate (GMO) and Chol in different ratios and determined their cytotoxicity and incorporation into cell membranes and/or cells by immunofluorescence staining. Our results show that these newly designed nanoparticles are not cytotoxic and that their incorporation into MDCK II cell membranes (used as a model system) may provide a mechanism that could be applied to RPE cells expressing mutated hBest1 in order to restore their ion transport functions, affected by mutated and malfunctioning hBest1 molecules.
Keywords: MDCK II cells; bilayer nanostructures; hBest1.
Conflict of interest statement
The authors declare no conflicts of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Figures
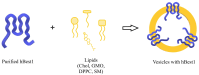
Similar articles
-
Cholesterol Alters the Phase Separation in Model Membranes Containing hBest1.Molecules. 2022 Jul 2;27(13):4267. doi: 10.3390/molecules27134267. Molecules. 2022. PMID: 35807512 Free PMC article.
-
Effects of Ca2+, Glu and GABA on hBest1 and composite hBest1/POPC surface films.Colloids Surf B Biointerfaces. 2018 Jan 1;161:192-199. doi: 10.1016/j.colsurfb.2017.10.051. Epub 2017 Oct 18. Colloids Surf B Biointerfaces. 2018. PMID: 29080503
-
Lipid membrane behavior of nitro-fatty acids and their loading into liposomes to activate Nrf2 pathway in RAW264.7 cells with impact on intracellular NO production.Chem Phys Lipids. 2025 Aug;270:105497. doi: 10.1016/j.chemphyslip.2025.105497. Epub 2025 May 16. Chem Phys Lipids. 2025. PMID: 40383219
-
Management of urinary stones by experts in stone disease (ESD 2025).Arch Ital Urol Androl. 2025 Jun 30;97(2):14085. doi: 10.4081/aiua.2025.14085. Epub 2025 Jun 30. Arch Ital Urol Androl. 2025. PMID: 40583613 Review.
-
Systemic treatments for metastatic cutaneous melanoma.Cochrane Database Syst Rev. 2018 Feb 6;2(2):CD011123. doi: 10.1002/14651858.CD011123.pub2. Cochrane Database Syst Rev. 2018. PMID: 29405038 Free PMC article.
References
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

