The OsAP4-OsCATA/OsCATC Regulatory Module Orchestrates Drought Stress Adaptation in Rice Seedlings Through ROS Scavenging
- PMID: 40733410
- PMCID: PMC12298810
- DOI: 10.3390/plants14142174
The OsAP4-OsCATA/OsCATC Regulatory Module Orchestrates Drought Stress Adaptation in Rice Seedlings Through ROS Scavenging
Abstract
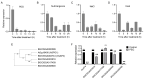
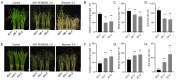
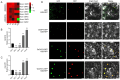
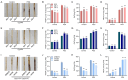
Drought stress poses a major constraint on global crop productivity. Although aspartic proteases (APs) are primarily characterized in plant disease resistance, their roles in abiotic stress adaptation remain largely unexplored. Here, we demonstrate that rice (Oryza sativa) OsAP4 critically regulates drought stress tolerance at the seedling stage. Genetic manipulation through overexpression (OsAP4-OE) or CRISPR knockout (OsAP4-KO) resulted in significantly reduced or enhanced stress tolerance compared to wild-type plants, respectively. Through integrated approaches including yeast two-hybrid, bimolecular fluorescence complementation, pull-down, co-immunoprecipitation, and protein degradation assays, we established that OsAP4 physically interacts with and destabilizes OsCATA/OsCATC, two catalase enzymes responsible for reactive oxygen species (ROS) scavenging. Importantly, OsAP4 modulates ROS production under drought stress treatment conditions. Together, these findings reveal a novel OsAP4-OsCATA/OsCATC regulatory module governing rice drought stress responses.
Keywords: OsAP4; OsCATs; ROS scavenging; drought stress; rice.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures

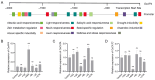
Similar articles
-
The OsFBX388-OsFIP1-OsCatA module regulates ROS homeostasis and disease resistance in rice.New Phytol. 2025 Sep;247(6):2871-2884. doi: 10.1111/nph.70383. Epub 2025 Jul 12. New Phytol. 2025. PMID: 40650454
-
OsCPK9-mediated Thr105 phosphorylation activates OsCATC to enhance drought tolerance in rice.J Adv Res. 2025 Jun 5:S2090-1232(25)00390-X. doi: 10.1016/j.jare.2025.06.004. Online ahead of print. J Adv Res. 2025. PMID: 40482964
-
OsCPK12 phosphorylates OsCATA and OsCATC to regulate H2O2 homeostasis and improve oxidative stress tolerance in rice.Plant Commun. 2024 Mar 11;5(3):100780. doi: 10.1016/j.xplc.2023.100780. Epub 2023 Dec 21. Plant Commun. 2024. PMID: 38130060 Free PMC article.
-
Epitranscriptomic Control of Drought Tolerance in Rice: The Role of RNA Methylation.Plants (Basel). 2025 Jun 30;14(13):2002. doi: 10.3390/plants14132002. Plants (Basel). 2025. PMID: 40648011 Free PMC article. Review.
-
Unveiling the secrets of abiotic stress tolerance in plants through molecular and hormonal insights.3 Biotech. 2024 Oct;14(10):252. doi: 10.1007/s13205-024-04083-7. Epub 2024 Sep 26. 3 Biotech. 2024. PMID: 39345964 Review.
References
-
- Mutlu A., Gal S. Plant aspartic proteinases: Enzymes On the way to a function. Physiol. Plant. 1999;105:569–576. doi: 10.1034/j.1399-3054.1999.105324.x. - DOI
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

