Genome-Wide Identification and Functional Characterization of the BAHD Acyltransferase Gene Family in Brassica napus L
- PMID: 40733420
- PMCID: PMC12299259
- DOI: 10.3390/plants14142183
Genome-Wide Identification and Functional Characterization of the BAHD Acyltransferase Gene Family in Brassica napus L
Abstract
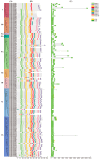
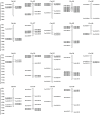
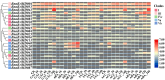
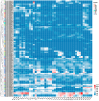
The BAHD acyltransferase family plays a critical role in plant secondary metabolism by catalyzing acyl transfer reactions that are essential for synthesizing metabolites involved in environmental adaptation. However, systematic investigation of this superfamily in Brassica napus has not been reported. In this study, 158 BnaBAHD genes were identified by comprehensive analyses of evolutionary relationships, motif structures, chromosomal distribution, gene collinearity, and selection pressures, and these genes were phylogenetically classified into five clades harboring conserved catalytic domains (HXXXD and DFGWG). Transient overexpression combined with metabolomic profiling demonstrated that two homologous seed-specific Clade V members, BnaBAHD040 and BnaBAHD120, which exhibited elevated expression during late seed development, significantly enhanced the accumulation of acylated metabolites contributing to biotic/abiotic stress resistance. This study provides the first experimental validation of the catalytic functions of BAHD enzymes in B. napus, establishing a theoretical foundation for leveraging this gene family in genetic improvement to develop novel rapeseed cultivars with enhanced stress tolerance and yield.
Keywords: BAHD acyltransferase; Brassica napus L.; acylation; stress resistance.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures





Similar articles
-
Genome-wide identification and evolutionary analysis of the NRAMP gene family in the AC genomes of Brassica species.BMC Plant Biol. 2024 Apr 23;24(1):311. doi: 10.1186/s12870-024-04981-1. BMC Plant Biol. 2024. PMID: 38649805 Free PMC article.
-
Evolutionary dynamics of the calcium/cation antiporter superfamily in Brassicaceae: codon usage, selection pressure, and BnCaCAs role in abiotic stress response.Front Plant Sci. 2025 Jul 8;16:1506461. doi: 10.3389/fpls.2025.1506461. eCollection 2025. Front Plant Sci. 2025. PMID: 40697863 Free PMC article.
-
Genome-wide identification and functional analysis of PEL gene family in Brassica napus L.BMC Plant Biol. 2025 Jul 2;25(1):838. doi: 10.1186/s12870-025-06864-5. BMC Plant Biol. 2025. PMID: 40604400 Free PMC article.
-
The Black Book of Psychotropic Dosing and Monitoring.Psychopharmacol Bull. 2024 Jul 8;54(3):8-59. Psychopharmacol Bull. 2024. PMID: 38993656 Free PMC article. Review.
-
[Volume and health outcomes: evidence from systematic reviews and from evaluation of Italian hospital data].Epidemiol Prev. 2013 Mar-Jun;37(2-3 Suppl 2):1-100. Epidemiol Prev. 2013. PMID: 23851286 Italian.
References
-
- Pascal S., Bernard A., Sorel M., Pervent M., Vile D., Haslam R.P., Napier J.A., Lessire R., Domergue F., Joubès J. The Arabidopsis Cer26 Mutant, like the Cer26 Mutant, Is Specifically Affected in the Very Long Chain Fatty Acid Elongation Process. Plant J. 2013;73:733–746. doi: 10.1111/tpj.12060. - DOI - PubMed
-
- Zhang W., Li J., Dong Y., Huang Y., Qi Y., Bai H., Li H., Shi L. Genome-Wide Identification and Expression of BAHD Acyltransferase Gene Family Shed Novel Insights into the Regulation of Linalyl Acetate and Lavandulyl Acetate in Lavender. J. Plant Physiol. 2024;292:154143. doi: 10.1016/j.jplph.2023.154143. - DOI - PubMed
-
- Xu L., Zeisler V., Schreiber L., Gao J., Hu K., Wen J., Yi B., Shen J., Ma C., Tu J., et al. Overexpression of the Novel Arabidopsis Gene At5g02890 Alters Inflorescence Stem Wax Composition and Affects Phytohormone Homeostasis. Front. Plant Sci. 2017;8:68. doi: 10.3389/fpls.2017.00068. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources

