Mitochondria-derived nuclear ATP surge protects against confinement-induced proliferation defects
- PMID: 40738970
- PMCID: PMC12310956
- DOI: 10.1038/s41467-025-61787-x
Mitochondria-derived nuclear ATP surge protects against confinement-induced proliferation defects
Abstract
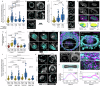
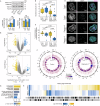
The physical tissue microenvironment regulates cell state and behaviour. How mechanical confinement rewires the subcellular localisation of organelles and affects cellular metabolism is largely unknown. In this study, proteomics analysis revealed that cellular confinement induced a strong enrichment of mitochondrial proteins in the nuclear fraction. Quantitative live cell microscopy confirmed that mechanical cell confinement leads to a rapid re-localisation of mitochondria to the nuclear periphery in vitro, reflecting a physiologically relevant phenomenon in patient-derived tumours. This nucleus-mitochondria proximity is mediated by an endoplasmic reticulum-based net that entraps the mitochondria in an actin-dependent manner. Functionally, the nucleus-mitochondria proximity results in a nuclear ATP surge, which can be regulated by the genetic and pharmacological modulation of mitochondrial ATP production or via alterations of the actin cytoskeleton. The confinement-induced nuclear ATP surge has physiologically significant long-term effects on cell fitness, driven by changes in chromatin state, enhanced DNA damage repair, and cell cycle progression during mechanical cell deformation. Together, our data describe a confinement-induced metabolic adaptation that is required to enable prompt DNA damage repair and cell proliferation under mechanical confinement stress by facilitating chromatin state transitions.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures




References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

