Human umbilical cord mesenchymal stem cells ameliorate liver fibrosis by inhibiting hepatocyte ferroptosis and macrophage polarization via the miR-455-3p/PLAU axis
- PMID: 40739265
- PMCID: PMC12312308
- DOI: 10.1186/s13287-025-04526-9
Human umbilical cord mesenchymal stem cells ameliorate liver fibrosis by inhibiting hepatocyte ferroptosis and macrophage polarization via the miR-455-3p/PLAU axis
Abstract
Background: Peripheral infusion of human umbilical cord mesenchymal stem cells (hUC-MSCs) may therapeutically treat liver fibrosis. hUC-MSCs can influence liver fibrosis progression and immune microenvironment remodeling via microRNAs (miRNAs).
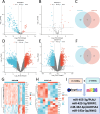
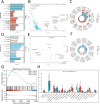
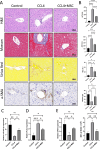
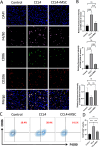
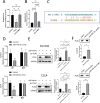
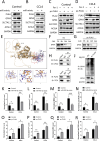
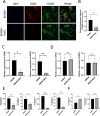
Methods: Bioinformatics showed the miR-455-3p/PLAU axis might regulate ferroptosis in liver fibrosis and affect the immune microenvironment. hUC-MSCs were transplanted into CCL4 induced liver-fibrotic male C57BL/6 mice. Liver tissues were stained (Hematoxylin and eosin (H&E), Masson, Sirius Red) to assess fibrosis; Alpha-smooth muscle actin (α-SMA) expression was detected by immunohistochemistry (IHC). Macrophage polarization was measured by immunofluorescence (IF), quantitative real-time polymerase chain reaction (qRT-PCR), and flow cytometry. Dual-luciferase and RNA pull-down assays verified miR-455-3p/PLAU regulation. Alanine aminotransferase (ALT), aspartate aminotransferase (AST), and total bilirubin (TBIL) evaluated liver function. Reactive oxygen species (ROS), glutathione peroxidase 4 (GPX4), iron content, and ferrous iron (Fe2+) assessed ferroptosis. Enzyme-linked immunosorbent assay (ELISA) detected cytokines in supernatant.
Results: In fibrotic vs. normal liver tissues, miR-455-3p was down-regulated, PLAU up-regulated, and ferroptosis increased. hUC-MSCs transplantation improves CCL4 induced liver fibrosis by inhibiting ferroptosis of hepatocytes through the miR-455-3p/PLAU axis. It also alleviated macrophage M1 polarization and liver damage.
Conclusion: hUC-MSCs inhibit hepatocyte ferroptosis via the miR-455-3p/PLAU axis, alleviate macrophage M1 polarization, and slow liver fibrosis. These findings offer insights for future liver fibrosis research and treatment.
Keywords: Ferroptosis; Human umbilical cord mesenchymal stem cells; Liver fibrosis; Macrophage polarization; MiR-455-3p/PLAU.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: Informed consent and approval were obtained from all the patients and the Ethics Committee of the Affiliated Hospital of Jiangsu University (Study on improvement of liver fibrosis by mesenchymal stem cells, KY2022K0103, Date: 2022-01-03). All the animal studies were performed following the Guidelines for the Care and Use of Laboratory Animals and were approved by the Ethics Committee of Jiangsu University (The role and application of mesenchymal stem cells in regulating ferroptosis in hepatic fibrosis, UJS-IACUC-AP-2024073031, Date: 2024-07-30). Consent for publication: Informed consent was obtained from all individual participants included in the study. Competing interests: The authors declare that they have no competing interests.
Figures
References
-
- Kisseleva T, Brenner D. Molecular and cellular mechanisms of liver fibrosis and its regression. Nat Rev Gastroenterol Hepatol. 2021;18(3):151–66. - PubMed
-
- Hernandez-Gea V, Friedman SL. Pathogenesis of liver fibrosis. Annu Rev Pathol. 2011;6:425–56. - PubMed
-
- Pimpin L, et al. Burden of liver disease in Europe: epidemiology and analysis of risk factors to identify prevention policies. J Hepatol. 2018;69(3):718–35. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

