Prediction of Small-Molecule Partitioning into Biomolecular Condensates from Simulation
- PMID: 40747072
- PMCID: PMC12308379
- DOI: 10.1021/jacsau.5c00291
Prediction of Small-Molecule Partitioning into Biomolecular Condensates from Simulation
Abstract
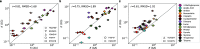
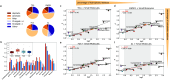
Predicting small-molecule partitioning into biomolecular condensates is the key to developing drugs that selectively target aberrant condensates. However, the molecular mechanisms underlying small-molecule partitioning remain largely unknown. Here, we first exploit atomistic molecular dynamics simulations of model condensates to elucidate the physicochemical rules governing small-molecule partitioning. We find that while hydrophobicity is a key factor in determining partitioning into condensates enriched in hydrophobic residues, partitioning into more polar condensates is driven by specific interactions that can offset the associated entropic cost of localization. The observed selectivity of condensates toward certain compounds suggests that condensate-specific therapeutics can be engineered. Building on these insights, we develop minimal models (MAPPS) for the efficient prediction of small-molecule partitioning into biologically relevant condensates. We demonstrate that this approach reproduces atomistic partition coefficients in both model systems and condensates composed of the low-complexity domain (LCD) of FUS. Applying MAPPS to various LCD-based condensates shows that the protein sequence can exert a selective pressure, thereby influencing small-molecule partitioning. Collectively, our findings reveal that partitioning is driven by both small molecule-protein affinity and the complex interplay between the physicochemical properties of the compounds and the condensate environment.
Keywords: biomolecular condensates; coarse-grained model; molecular dynamics; partitioning; small molecules.
© 2025 The Authors. Published by American Chemical Society.
Figures
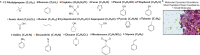



Update of
-
Prediction of small-molecule partitioning into biomolecular condensates from simulation.bioRxiv [Preprint]. 2025 Mar 6:2025.03.04.641530. doi: 10.1101/2025.03.04.641530. bioRxiv. 2025. Update in: JACS Au. 2025 Jul 03;5(7):3125-3139. doi: 10.1021/jacsau.5c00291. PMID: 40093099 Free PMC article. Updated. Preprint.
References
Grants and funding
LinkOut - more resources
Full Text Sources
