A multicellular self-organized probiotic platform for oral delivery enhances intestinal colonization
- PMID: 40750778
- PMCID: PMC12317005
- DOI: 10.1038/s41467-025-62349-x
A multicellular self-organized probiotic platform for oral delivery enhances intestinal colonization
Abstract
Stable gut colonization of probiotics is essential for sustained therapeutic effects, however traditional oral probiotic supplements often fail to adapt to the gut environment. Here, based on the observation that multicellular microcolonies instead of planktonic bacteria display a more advantageous gene pattern for colonization, we design a system encapsulating multicellular self-organized probiotic microcolonies, termed Express Microcolony Service (EMS), for efficient oral delivery and enhanced gut colonization of probiotics. Utilizing the stress-relaxing and acid-resistant property of the covalent-ionic crosslinking alginate hydrogel microsphere, the micro-cargo provides tunable nutrient supply and extracellular matrix support to facilitate microcolony self-organization. Notably, we show that the variable spatial constraints of the stress-relaxing hydrogel could modulate the viability and colonization potential of microcolonies. In vitro, bacteria microcolonies in EMS show remarkable resistance to gastric acid, bile salts, and antibiotics, compared with planktonic probiotics. In vivo, the EMS strategy exhibits 89- and 52-fold higher colonization rate in the cecum and colon of mice, compared to conventional oral probiotics. The multicellular self-organized EMS system offers an innovative, efficient and clinically transformable alternative for probiotic therapy.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures

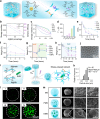




References
-
- O’Toole, P. W., Marchesi, J. R. & Hill, C. Next-generation probiotics: the spectrum from probiotics to live biotherapeutics. Nat. Microbiol.2, 17057 (2017). - PubMed
-
- Li, N. et al. The role of gut microbiota associated metabolites in digestive disorders. Eng. Regener.5, 228–246 (2024).
-
- Veiga, P., Suez, J., Derrien, M. & Elinav, E. Moving from probiotics to precision probiotics. Nat. Microbiol.5, 878–880 (2020). - PubMed
-
- Guyonnet, D., Schlumberger, A., Mhamdi, L., Jakob, S. & Chassany, O. Fermented milk containing Bifidobacterium lactis DN-173 010 improves gastrointestinal well-being and digestive symptoms in women reporting minor digestive symptoms: a randomised, double-blind, parallel, controlled study. Br. J. Nutr.102, 1654–1662 (2009). - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

