Analyzing Shigella in Wuhan: Serotypes, Antimicrobial Resistance, and Public Health Implications
- PMID: 40756687
- PMCID: PMC12316178
- DOI: 10.2147/IDR.S522808
Analyzing Shigella in Wuhan: Serotypes, Antimicrobial Resistance, and Public Health Implications
Abstract
Objective: This study aims to delineate the epidemiological characteristics and antibiotic resistance of Shigella flexneri isolates from Wuhan, focusing on serotype distribution, resistance patterns, and genetic diversity.
Methods: Our study analyzed 40Shigella flexneri isolates collected from 2011 to 2022 in Wuhan, assessing their serotype distribution and resistance to multiple antibiotics. We conducted resistance gene detection and genetic diversity analysis using polymerase chain reaction and pulsed-field gel electrophoresis (PFGE), respectively.
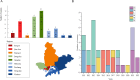
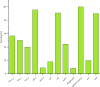
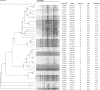
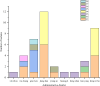
Results: The study revealed significant clustering of S. flexneri in the Jianghan and Dongxihu districts, with serotype 2a predominating. Isolates exhibited high resistance to ampicillin, tetracycline, and trimethoprim-sulfamethoxazole, with an overall multidrug resistance (MDR) rate of 67.5%. Serotypes 1b and 2b were fully sensitive, contrasting with higher resistance in serotypes 2a and 4a to fluoroquinolones. Resistance mechanisms included blaOXA and blaTEM for ampicillin, blaCTX-M for cephalosporins, tetB for tetracycline, and dfrA1 and sul2 for trimethoprim-sulfamethoxazole. All 12 quinolone-resistant isolates exhibited mutations in gyrA (S83L, D87N, D87G), parC (S80I), and parE (458A), and novel mutations were identified in gyrA (H221Y, A221V, A221E, I222N), parC (D197A), and parE (G408D). Pulsed-field gel electrophoresis (PFGE) analysis highlighted extensive genetic diversity with dominant groups P1 and P4, and with notable regional and temporal distribution patterns. Distinct PFGE types exhibited unique antimicrobial resistance profiles, with P1 and P4 showing high rates of multidrug resistance, while P5 and P3 displayed lower resistance levels. A notable evolutionary adaptation was observed in a clone from 2016 (P4-1), which by 2017 (P4-2) had acquired aminoglycoside resistance.
Conclusion: The study underscores the significant regional specificity and genetic diversity of S. flexneri in Wuhan, which poses challenges for treatment due to high antibiotic resistance and MDR prevalence. Findings stress the need for enhanced surveillance and tailored public health strategies to manage shigellosis effectively.
Keywords: Shigella flexneri; epidemiology; multidrug resistance; pulsed-field gel electrophoresis; resistance profile.
© 2025 Deng et al.
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures
Similar articles
-
Comprehensive analysis of extensive drug-resistant Salmonella Typhi in Gujarat region, India: genomic findings and prospective alternative therapy.Microbiol Spectr. 2025 Jul;13(7):e0254024. doi: 10.1128/spectrum.02540-24. Epub 2025 May 27. Microbiol Spectr. 2025. PMID: 40422861 Free PMC article.
-
Pan-drug, colistin, streptomycin, erythromycin, clindamycin resistant Salmonella enterica serovars isolated from slaughtered cattle and human in mansoura, Egypt.Ann Clin Microbiol Antimicrob. 2025 Jul 3;24(1):40. doi: 10.1186/s12941-025-00809-4. Ann Clin Microbiol Antimicrob. 2025. PMID: 40611300 Free PMC article.
-
Antibiotic-Resistance Profiles and Genetic Diversity of Shigella Isolates in China: Implications for Control Strategies.Foodborne Pathog Dis. 2024 Jun;21(6):378-385. doi: 10.1089/fpd.2023.0138. Epub 2024 Apr 1. Foodborne Pathog Dis. 2024. PMID: 38557159
-
Prophylactic antibiotics for adults with chronic obstructive pulmonary disease: a network meta-analysis.Cochrane Database Syst Rev. 2021 Jan 15;1(1):CD013198. doi: 10.1002/14651858.CD013198.pub2. Cochrane Database Syst Rev. 2021. PMID: 33448349 Free PMC article.
-
A systematic review and meta-analysis revealed a high-level antibiotic resistance of bovine mastitis Staphylococcus aureus in Iran.Res Vet Sci. 2023 Aug;161:23-30. doi: 10.1016/j.rvsc.2023.05.016. Epub 2023 Jun 5. Res Vet Sci. 2023. PMID: 37302281
References
-
- Moxley RA. Enterobacteriaceae: shigella. Veterinary Microbiol. 2022;100–107.
LinkOut - more resources
Full Text Sources
Miscellaneous

