Artificial intelligence and machine learning in the development of vaccines and immunotherapeutics-yesterday, today, and tomorrow
- PMID: 40756816
- PMCID: PMC12313644
- DOI: 10.3389/frai.2025.1620572
Artificial intelligence and machine learning in the development of vaccines and immunotherapeutics-yesterday, today, and tomorrow
Abstract
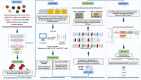
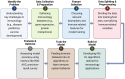
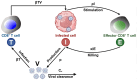
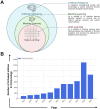
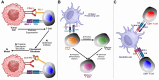
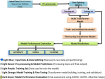
The development of vaccines and immunotherapies against infectious diseases and cancers has been one of the significant achievements of medical science in the last century. Subunit vaccines offer key advantages over whole-inactivated or attenuated-pathogen-based vaccines, as they elicit more specific B-and T-cell responses with improved safety, immunogenicity, and protective efficacy. However, developing subunit vaccines is often cost-and time-consuming. In the past, the development of vaccines and immunotherapeutics relied heavily on trial-and-error experimentation, as well as extensive and costly in vivo testing, which typically required years of pre-clinical and clinical trials. Today, artificial intelligence (AI) and deep learning (DL) are actively transforming vaccine and immunotherapeutic research by (i) offering predictive frameworks that support rapid, data-driven decision-making, (ii) integrating computational models, systems vaccinology, and multi-omics data (iii) helping to better phenotype, differentiate, and classify patients diseases and cancers; (iv), integrating host characteristics for tailored vaccines and immunotherapeutics; (v) refining the selection of B-and T-cell antigen/epitope targets to enhance efficacy and durability of immune protection; and (vi) enabling a deeper understanding of immune regulation, immune evasion, and regulatory pathways. Artificial intelligence and DL are pushing the boundaries toward (i) the potential replacement of animal preclinical testing of vaccines and immunotherapeutics with computational-based models, as recently proposed by the United States NIH and FDA, and (ii) improving clinical trials by enabling real-time modeling for immune-bridging, predicting patients' immune responses, safety, and protective efficacy to vaccines and immunotherapeutics. In this review, we describe the past and current applications of AI and DL as time-and resource-efficient strategies and discuss future challenges in implementing AI and DL as new transformative fields that may facilitate the rapid development of precision and personalized vaccines and immunotherapeutics for infectious diseases and cancers.
Keywords: artificial intelligence (AI); cancer; immunotherapeutics; infectious diseases; machine learning (ML); vaccine development.
Copyright © 2025 Elfatimi, Lekbach, Prakash, and BenMohamed.
Conflict of interest statement
LB has an equity interest in TechImmune, LLC, a company that may potentially benefit from the research results and serves on the company’s Scientific Advisory Board. LB’s relationship with TechImmune, LLC, has been reviewed and approved by the University of California, Irvine, in accordance with its conflict-of-interest policies. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The author(s) declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
Figures

References
-
- Anderson L. N., Hoyt C. T., Zucker J. D., McNaughton A. D., Teuton J. R., Karis K., et al. (2025). Computational tools and data integration to accelerate vaccine development: challenges, opportunities, and future directions. Front. Immunol. 16:1502484. doi: 10.3389/fimmu.2025.1502484, PMID: - DOI - PMC - PubMed
-
- Bhinder B., Gilvary C., Madhukar N. S., Elemento O. (2021). Artificial intelligence in Cancer research and precision medicine. Cancer Discov. 11, 900–915. doi: 10.1158/2159-8290.CD-21-0090, PMID: - DOI - PMC - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Miscellaneous

