Rational engineering of allosteric protein switches by in silico prediction of domain insertion sites
- PMID: 40759748
- PMCID: PMC12328240
- DOI: 10.1038/s41592-025-02741-z
Rational engineering of allosteric protein switches by in silico prediction of domain insertion sites
Abstract
Domain insertion engineering is a powerful approach to juxtapose otherwise separate biological functions, resulting in proteins with new-to-nature activities. A prominent example are switchable protein variants, created by receptor domain insertion into effector proteins. Identifying suitable, allosteric sites for domain insertion, however, typically requires extensive screening and optimization. We present ProDomino, a machine learning pipeline to rationalize domain recombination, trained on a semisynthetic protein sequence dataset derived from naturally occurring intradomain insertion events. ProDomino robustly identifies domain insertion sites in proteins of biotechnological relevance, which we experimentally validated in Escherichia coli and human cells. Finally, we used light- and chemically regulated receptor domains as inserts and demonstrate the rapid, model-guided creation of potent, single-component opto- and chemogenetic protein switches. These include novel CRISPR-Cas9 and -Cas12a variants for inducible genome engineering in human cells. Our work enables one-shot domain insertion engineering and substantially accelerates the design of customized allosteric proteins.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures




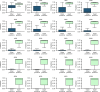

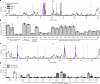


References
-
- Ponting, C. P. & Russell, R. R. The natural history of protein domains. Annu. Rev. Biophys. Biomol. Struct.31, 45–71 (2002). - PubMed
-
- Apic, G. & Russell, R. B. Domain recombination: a workhorse for evolutionary innovation. Sci. Signal.3, 139 (2010). - PubMed
-
- Mathony, J. & Niopek, D. Enlightening allostery: designing switchable proteins by photoreceptor fusion. Adv. Biol.5, 2000181 (2021). - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

