Blocking CXCR4+ CD4+ T cells reprograms Treg-mediated immunosuppression via modulating the Rho-GTPase/NF-κB signaling axis
- PMID: 40760450
- PMCID: PMC12323159
- DOI: 10.1186/s13073-025-01515-8
Blocking CXCR4+ CD4+ T cells reprograms Treg-mediated immunosuppression via modulating the Rho-GTPase/NF-κB signaling axis
Abstract
Background: While clinical trials have shown that CXCR4 antagonists can enhance the efficacy of cancer immunotherapy, the molecular mechanisms by which CXCR4 modulates the tumor microenvironment remain poorly understood. We recently identified CXCR4 as a regulator of exhausted CD8+ T cell phenotypes in cancer. Here, we investigate its role in orchestrating regulatory T (Treg) cell-mediated immunosuppression within tumors.
Methods: We conducted meta-analyses of single-cell RNA-seq datasets from pan-cancer tissues to characterize CXCR4 expression patterns in CD4+ T cells. Using CXCR4 antagonists and conditional knockout mice (Cxcr4flox/flox, LckCre), we inhibited Treg phenotypes in vivo. Through single-cell transcriptomics and single-cell ATAC-seq of the cervical cancer mouse model, phosphoproteomics, and ChIP-seq analyses, we elucidated how CXCR4 blockade in CD4+ T cells suppresses activated Treg phenotypes by modulating the Rho-GTPase/NF-κB signaling axis. We further integrated RNA-seq data, clinical trial datasets (NCT02826486 and NCT04516616), and human organoid models to validate the therapeutic potential of CXCR4 inhibition in enhancing antitumor immunotherapy.
Results: Single-cell transcriptomics of CD4+ T cells across multiple cancers revealed CXCR4 expression was associated with Treg cell developmental trajectories. Pharmacological and genetic inhibition of CXCR4 inhibited Treg phenotypes in cervical cancer and breast cancer. Mechanistically, phosphoproteomics and ChIP-seq analyses unveiled that blocking CXCR4+ CD4+ T cells reduced activated Treg phenotypes by modulating the Rho-GTPase/NF-κB signaling axis. Single-cell transcriptomic and multi-omic analyses demonstrated that blocking CXCR4+ CD4+ T cells promoted immunotherapy via reprogramming Treg-mediated immunosuppression. Furthermore, clinical trial data and human cervical cancer organoids confirmed that blocking CXCR4 enhances antitumor immunotherapy by reducing Treg phenotypes.
Conclusions: Our study highlights the crucial role of CXCR4 in deriving Treg-mediated immunosuppression via regulating the Rho-GTPase/NF-κB signaling axis, informing the potential of combining CXCR4 blockades with T cell-targeted immunotherapies.
Keywords: CXCR4; Immunosuppression; NF-κB2; Regulatory T (Treg) cells; Rho GTPase pathway.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: This study was approved by the Ethics and Institutional Review Board of Tongji Hospital, Tongji Medical College, Huazhong University of Science and Technology, China (Approval No. TJ-IRB20230855). A waiver of written informed consent was granted because the patient-derived biospecimens used in this study were obtained from surplus surgical samples that would otherwise be discarded. Clinical specimens were registered and obtained from the Biological Specimen Bank of Tongji Hospital in full compliance with the principles of the Declaration of Helsinki. All specimens had been fully de-identified prior to their release for research use, ensuring the protection of patient privacy. The study involved no direct patient contact and posed minimal risk. Therefore, the IRB approved the waiver in accordance with institutional regulations and the Declaration of Helsinki. All animal experiments were reviewed and approved by the Institutional Animal Care and Use Committee (IACUC) of Tongji Hospital, Tongji Medical College, Huazhong University of Science and Technology (Approval No. TJH-20211109). All animal care and experimental procedures were conducted in accordance with the institutional guidelines, the ARRIVE guidelines, and the national regulations for the administration of laboratory animals in China (Regulations for the Administration of Affairs Concerning Experimental Animals, revised in 2017). Animal welfare was monitored throughout the study, and humane endpoints were applied where necessary. Euthanasia was performed in accordance with the AVMA Guidelines for the Euthanasia of Animals (2020 edition). Consent for publication: Not applicable. Competing interests: The authors declare no competing interests.
Figures






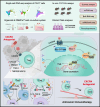
References
-
- Bockorny B, Semenisty V, Macarulla T, Borazanci E, Wolpin BM, Stemmer SM, et al. BL-8040, a CXCR4 antagonist, in combination with pembrolizumab and chemotherapy for pancreatic cancer: the COMBAT trial. Nat Med. 2020;26(6):878–85. - PubMed
-
- Choueiri TK, Atkins MB, Rose TL, Alter RS, Ju Y, Niland K, et al. A phase 1b trial of the CXCR4 inhibitor mavorixafor and nivolumab in advanced renal cell carcinoma patients with no prior response to nivolumab monotherapy. Invest New Drugs. 2021;39(4):1019–27. - PubMed
-
- Togashi Y, Shitara K, Nishikawa H. Regulatory T cells in cancer immunosuppression - implications for anticancer therapy. Nat Rev Clin Oncol. 2019;16(6):356–71. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

