High-throughput cultivation and isolation of environmental anaerobes using selectively permeable hydrogel capsules
- PMID: 40761568
- PMCID: PMC12319321
- DOI: 10.1093/ismeco/ycaf117
High-throughput cultivation and isolation of environmental anaerobes using selectively permeable hydrogel capsules
Abstract
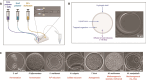
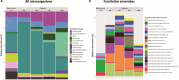
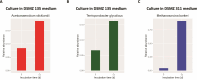
Over the past two decades, metagenomics has greatly expanded our understanding of microbial phylogenetic and metabolic diversity. However, most microbial taxa remain uncultured, hindering research and biotechnological applications. Isolating environmental anaerobes using traditional methods is particularly cumbersome and low throughput. Here, we present a novel, high-throughput approach for the cultivation and isolation of anaerobes, which involves trapping and growing single microbes within selectively permeable hydrogel capsules followed by fluorescence-activated cell sorting to distribute compartmentalized isolates into liquid medium for further growth. We show that diverse anaerobes can grow within capsules and that slower-growing ones (e.g. methanogens) can be enriched with this platform. We also applied this approach to isolate anaerobes from soil, including strains of the sulfate-reducing bacteria Desulfovibrio desulfuricans and Nitratidesulfovibrio vulgaris. Overall, this work introduces a robust, high-throughput alternative to traditional techniques for isolating environmental anaerobes and expands the emerging set of microfluidics-based tools for the cultivation of novel taxa.
Keywords: acetogenic bacteria; anaerobic microorganisms; hydrogel capsules; isolation; methanogens; microfluidics; soil microbiome; sulfate-reducing bacteria.
© The Author(s) 2025. Published by Oxford University Press on behalf of the International Society for Microbial Ecology.
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
Ultrahigh-throughput screening of environmental bacteria for proteolytic activity using droplet-based microfluidics.Appl Environ Microbiol. 2025 Jul 23;91(7):e0010925. doi: 10.1128/aem.00109-25. Epub 2025 Jun 13. Appl Environ Microbiol. 2025. PMID: 40511932 Free PMC article.
-
Culturing the uncultured anaerobes from the perspective of microbial growth curves.Trends Microbiol. 2025 Jul 3:S0966-842X(25)00182-9. doi: 10.1016/j.tim.2025.06.006. Online ahead of print. Trends Microbiol. 2025. PMID: 40615288 Review.
-
High-throughput library transgenesis in Caenorhabditis elegans via Transgenic Arrays Resulting in Diversity of Integrated Sequences (TARDIS).Elife. 2023 Jul 4;12:RP84831. doi: 10.7554/eLife.84831. Elife. 2023. PMID: 37401921 Free PMC article.
-
Short-Term Memory Impairment.2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 31424720 Free Books & Documents.
-
Management of urinary stones by experts in stone disease (ESD 2025).Arch Ital Urol Androl. 2025 Jun 30;97(2):14085. doi: 10.4081/aiua.2025.14085. Epub 2025 Jun 30. Arch Ital Urol Androl. 2025. PMID: 40583613 Review.
References
-
- BacDive (DSMZ) dashboard, https://bacdive.dsmz.de/dashboard (9 November 2024, date last accessed).
-
- LPSN (DSMZ) numbers, https://lpsn.dsmz.de/text/numbers (9 November 2024, date last accessed).
LinkOut - more resources
Full Text Sources
Miscellaneous

