PgtE protease enables virulent Salmonella to evade C3-mediated serum and neutrophil killing
- PMID: 40767554
- PMCID: PMC12345235
- DOI: 10.1128/mbio.03802-24
PgtE protease enables virulent Salmonella to evade C3-mediated serum and neutrophil killing
Abstract
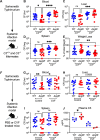
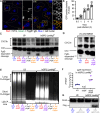
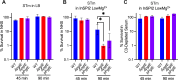
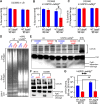
Non-typhoidal Salmonella serovars, such as Salmonella enterica serovar Typhimurium (STm), are a leading cause of inflammatory diarrhea in otherwise healthy individuals. Among children, the elderly, and immunocompromised individuals, STm can spread to systemic sites and cause potentially lethal bacteremia. Phagocytic cells and the immune complement system are pivotal to preventing the dissemination of STm. PgtE, an STm outer membrane protease, has been previously described to cleave over a dozen mammalian protein substrates in vitro, including complement protein C3. However, these activities have mostly been observed with mutant, avirulent strains with a truncated O-antigen that renders bacteria sensitive to complement killing. Here, we report that virulent STm utilizes PgtE to evade complement-mediated killing in vivo. The wild-type pathogen increases pgtE expression and PgtE proteolytic function within macrophages and in macrophage-like in vitro growth conditions, concomitant with physiologic O-antigen truncation in these environments. Furthermore, we found that wild-type STm's resistance to complement-mediated serum and neutrophil killing is PgtE-dependent. We propose that PgtE promotes the systemic spread of STm by acting as a second line of defense against complement when STm escapes from a macrophage.IMPORTANCENon-typhoidal Salmonella serovars primarily cause gastrointestinal infections but can also lead to bacteremia through mechanisms that are not completely elucidated. Here we show that the outer membrane protease PgtE enables virulent Salmonella to evade complement-mediated killing in vivo, thereby promoting bacteremia. We also demonstrate that pgtE expression and PgtE proteolytic function are increased within macrophages and under macrophage-like growth conditions. Upon escaping from macrophages, PgtE protects against complement-mediated killing by serum and neutrophils, promoting the systemic spread of the pathogen. These findings highlight PgtE as a potential therapeutic target for preventing Salmonella bacteremia.
Keywords: Salmonella; bacteremia; complement; macrophages; neutrophils; proteases.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Update of
-
PgtE protease enables virulent Salmonella to evade C3-mediated serum and neutrophil killing.bioRxiv [Preprint]. 2024 Nov 5:2024.11.05.622138. doi: 10.1101/2024.11.05.622138. bioRxiv. 2024. Update in: mBio. 2025 Aug 13;16(8):e0380224. doi: 10.1128/mbio.03802-24. PMID: 39574608 Free PMC article. Updated. Preprint.
References
-
- Majowicz SE, Musto J, Scallan E, Angulo FJ, Kirk M, O’Brien SJ, Jones TF, Fazil A, Hoekstra RM, International Collaboration on Enteric Disease “Burden of Illness” Studies . 2010. The global burden of nontyphoidal Salmonella gastroenteritis. Clin Infect Dis 50:882–889. doi: 10.1086/650733 - DOI - PubMed
MeSH terms
Substances
Grants and funding
- T32AI007036/National Institute of Allergy and Infectious Diseases
- AI145325/National Institute of Allergy and Infectious Diseases
- R01 AI129992/AI/NIAID NIH HHS/United States
- F32 AI169989/AI/NIAID NIH HHS/United States
- R01 AI145325/AI/NIAID NIH HHS/United States
- G-1019128.01/Burroughs Wellcome Fund
- T32 DK007202/DK/NIDDK NIH HHS/United States
- AI129992/National Institute of Allergy and Infectious Diseases
- JP233fa627003/Japan Agency for Medical Research and Development
- DK007202/DK/NIDDK NIH HHS/United States
- AI169989/National Institute of Allergy and Infectious Diseases
- T32 AI007036/AI/NIAID NIH HHS/United States
- MIST/National Institute of Allergy and Infectious Diseases
LinkOut - more resources
Full Text Sources
Miscellaneous
