Proteomic alterations in ovarian cancer-Predicting residual disease status using artificial intelligence and SHAP-based biomarker interpretation
- PMID: 40771481
- PMCID: PMC12325313
- DOI: 10.3389/fmed.2025.1562558
Proteomic alterations in ovarian cancer-Predicting residual disease status using artificial intelligence and SHAP-based biomarker interpretation
Abstract
Introduction: High-grade serous ovarian cancer (HGSOC) is the most aggressive and prevalent subtype of ovarian Treatment outcomes are significantly influenced by residual disease status following neoadjuvant chemotherapy (NACT). Predicting residual disease before surgery can improve patient stratification and personalized treatment strategies.
Methods: This study analyzed pre-NACT proteomic data from 20 HGSOC patients treated with NACT. Patients were categorized into two groups based on surgical outcomes: no residual disease (R0, n = 14) and suboptimal residual disease (R1, n = 6). From an initial set of 97 differentially expressed proteins, 18 significant proteins were selected using the BORUTA feature selection method. Three machine learning models-Random Forest (RF), Support Vector Machine (SVM), and Bootstrap Aggregation with Classification and Regression Trees (BaggedCART)-were developed and evaluated.
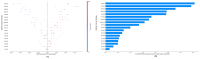
Results: The Random Forest model achieved the best performance with an AUC of 0.955, accuracy of 0.830, sensitivity of 0.904, specificity of 0.763, and F1-score of 0.839. SHapley Additive exPlanations (SHAP) analysis identified five proteins (P48637, O43491, O95302, Q96CX2, and P49189) as the most influential predictors of residual disease. These proteins, including glutathione synthetase and peptidyl-prolyl cis-trans isomerase FKBP9, are associated with chemotherapy resistance mechanisms.
Discussion: The findings demonstrate the potential of integrating proteomic data with machine learning techniques for predicting surgical outcomes in HGSOC. Identified protein signatures may serve as valuable biomarkers for anticipating NACT response and informing clinical decision-making, ultimately contributing to personalized patient care.
Keywords: SHAP analysis; high-grade serous ovarian cancer (HGSOC); machine learning; neoadjuvant chemotherapy (NACT); proteomic biomarkers.
Copyright © 2025 Yasar and Melekoglu.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
Similar articles
-
Impact of residual disease as a prognostic factor for survival in women with advanced epithelial ovarian cancer after primary surgery.Cochrane Database Syst Rev. 2022 Sep 26;9(9):CD015048. doi: 10.1002/14651858.CD015048.pub2. Cochrane Database Syst Rev. 2022. PMID: 36161421 Free PMC article.
-
Supervised Machine Learning Models for Predicting Sepsis-Associated Liver Injury in Patients With Sepsis: Development and Validation Study Based on a Multicenter Cohort Study.J Med Internet Res. 2025 May 26;27:e66733. doi: 10.2196/66733. J Med Internet Res. 2025. PMID: 40418571 Free PMC article.
-
Machine learning and SHAP value interpretation for predicting the response to neoadjuvant chemotherapy and long-term clinical outcomes in Chinese female breast cancer.Ann Med. 2025 Dec;57(1):2541316. doi: 10.1080/07853890.2025.2541316. Epub 2025 Aug 3. Ann Med. 2025. PMID: 40754951 Free PMC article.
-
Neoadjuvant chemotherapy before surgery versus surgery followed by chemotherapy for initial treatment in advanced epithelial ovarian cancer.Cochrane Database Syst Rev. 2025 Feb 10;2(2):CD005343. doi: 10.1002/14651858.CD005343.pub7. Cochrane Database Syst Rev. 2025. PMID: 39927569
-
Interpretable Machine Learning for Serum-Based Metabolomics in Breast Cancer Diagnostics: Insights from Multi-Objective Feature Selection-Driven LightGBM-SHAP Models.Medicina (Kaunas). 2025 Jun 19;61(6):1112. doi: 10.3390/medicina61061112. Medicina (Kaunas). 2025. PMID: 40572800 Free PMC article.
References
-
- Petrucelli N, Daly MB, Pal T. BRCA1- and BRCA2-associated hereditary breast and ovarian cancer. In:Adam MP, Feldman J, Mirzaa GM, Pagon RA, Wallace SE, Amemiya A, editors. GeneReviews®. Seattle, WA: University of Washington; (1998). p. 1993–2025. - PubMed
LinkOut - more resources
Full Text Sources

