O-GlcNAc modulation of nuclear pore complexes orchestrates mRNA export efficiency
- PMID: 40773237
- PMCID: PMC12358868
- DOI: 10.1073/pnas.2502687122
O-GlcNAc modulation of nuclear pore complexes orchestrates mRNA export efficiency
Abstract
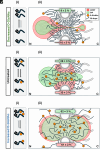
Efficient gene expression depends on the tightly regulated export of messenger RNA (mRNA) through nuclear pore complexes (NPCs), which are densely modified by O-linked N-acetylglucosamine (O-GlcNAc). Although dysregulated O-GlcNAcylation has been linked to a variety of human diseases, the precise distribution of O-GlcNAc within the NPC and its effects on mRNA export remain poorly understood. Here, we combined single-point edge-excitation subdiffraction (SPEED) microscopy with stochastic optical reconstruction microscopy (STORM) to map the nanometer-scale distribution of an O-GlcNAc analog (GlcNAz) within NPCs and to quantify the export kinetics of mRNA-protein complexes (mRNPs) under both normal and perturbed O-GlcNAcylation conditions. Under basal conditions, GlcNAz is predominantly localized around the central channel of the NPC. However, both hypo- and hyper-O-GlcNAcylation cause GlcNAz to redistribute toward the nuclear and cytoplasmic peripheries. This shift is paralleled by changes in mRNP localization and altered distributions of key, highly O-GlcNAcylated, phenylalanine-glycine nucleoporins. These architectural rearrangements are accompanied by functional consequences: Elevated O-GlcNAcylation nearly doubles mRNA export efficiency (~61%), while reduced O-GlcNAcylation lowers it to ~16%, along with reduced NPC engagement. The transport receptor TAP exhibits analogous efficiency changes, reinforcing the role of O-GlcNAcylation as a key regulator of nucleocytoplasmic transport. Together, these results suggest that O-GlcNAcylation modulates NPC architecture and transport dynamics to fine-tune mRNA export, and indicate that targeted modulation of NPC O-GlcNAc levels may offer a promising strategy for addressing diseases associated with nuclear transport dysfunction.
Keywords: O-GlcNAc; messenger RNA; nuclear pore complex; posttranslational modification; superresolution fluorescence microscopy.
Conflict of interest statement
Competing interests statement:The authors declare no competing interest.
Figures



References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

