SRS2 is required for MUS81-dependent CO formation in zmm mutants
- PMID: 40773506
- PMCID: PMC12349706
- DOI: 10.1371/journal.pgen.1011637
SRS2 is required for MUS81-dependent CO formation in zmm mutants
Abstract
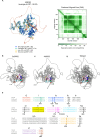
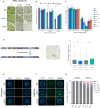
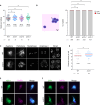
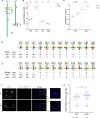
Helicases are enzymes that use the energy derived from ATP hydrolysis to translocate along and unwind nucleic acids. Accordingly, helicases are instrumental in maintaining genomic integrity and ensuring genetic diversity. Srs2 is a multi-functional DNA helicase that dismantles Rad51 nucleofilaments and regulates DNA strand invasion to prevent excessive or inappropriate homologous recombination in yeast. Consistently, the deletion of Srs2 has significant consequences for the maintenance of genome integrity in mitotic cells. In contrast, its role in meiotic recombination remains less clear. We present here substantial evidence that SRS2 plays an important role in meiotic recombination in the model plant Arabidopsis thaliana. Arabidopsis srs2 mutants exhibit moderate defects in DNA damage-induced RAD51 focus formation, but SRS2 is dispensable for DNA repair and RAD51-dependent recombination in somatic cells. Meiotic progression and fertility appear unaffected in srs2 plants but, strikingly, the absence of SRS2 leads to increased genetic interference accompanied by increased numbers of Class I COs and a reduction in MUS81-dependent Class II COs. We propose that SRS2 plays a role in MUS81-mediated resolution of a subset of recombination intermediates into Class II CO. The absence of SRS2 would thus lead to the alternative channeling of these recombination intermediates into the Class I CO pathway, resulting in an increased proportion of Class I CO.
Copyright: © 2025 Petiot et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
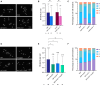

Similar articles
-
A large C-terminal Rad52 segment acts as a chaperone to Form and Stabilize Rad51 Filaments.Nat Commun. 2025 Jul 1;16(1):5589. doi: 10.1038/s41467-025-60664-x. Nat Commun. 2025. PMID: 40595518 Free PMC article.
-
The Main Role of Srs2 in DNA Repair Depends on Its Helicase Activity, Rather than on Its Interactions with PCNA or Rad51.mBio. 2018 Jul 17;9(4):e01192-18. doi: 10.1128/mBio.01192-18. mBio. 2018. PMID: 30018112 Free PMC article.
-
Prescription of Controlled Substances: Benefits and Risks.2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 30726003 Free Books & Documents.
-
Overexpression of RAD51 suppresses recombination defects: a possible mechanism to reverse genomic instability.Nucleic Acids Res. 2010 Mar;38(4):1061-70. doi: 10.1093/nar/gkp1063. Epub 2009 Nov 26. Nucleic Acids Res. 2010. PMID: 19942681 Free PMC article. Review.
-
Positive and negative regulators of RAD51/DMC1 in homologous recombination and DNA replication.DNA Repair (Amst). 2024 Feb;134:103613. doi: 10.1016/j.dnarep.2023.103613. Epub 2023 Dec 13. DNA Repair (Amst). 2024. PMID: 38142595 Review.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

