Integrative multiomics analysis reveals the subtypes and key mechanisms of platinum resistance in gastric cancer: identification of KLF9 as a promising therapeutic target
- PMID: 40775648
- PMCID: PMC12330134
- DOI: 10.1186/s12967-025-06725-7
Integrative multiomics analysis reveals the subtypes and key mechanisms of platinum resistance in gastric cancer: identification of KLF9 as a promising therapeutic target
Abstract
Background: Gastric cancer (GC) is characterized by significant intertumoral heterogeneity, which often leads to the development of resistance to platinum-based chemotherapy. Combining platinum drugs with other therapeutic strategies may improve treatment efficacy; however, the mechanisms underlying platinum resistance in GC remain unclear.
Methods: Key genes related to platinum resistance in GC were selected from the platinum resistance gene database and GC resistance datasets. The Similarity Network Fusion (SNF) algorithm was employed, along with prognosis-related methylation data and somatic mutation data, to classify the molecular subtypes of GC based on GC platinum resistance genes. Gene expression profiles, prognosis, immune cell infiltration, chemotherapy sensitivity, and immunotherapy responsiveness were comprehensively evaluated for each subtype. Localization and functional evaluation were conducted at the single-cell and spatial transcriptomics levels, and predictive models were developed using machine learning techniques. These functional differences in platinum resistance gene models were further explored in GC. Moreover, experimental validation was conducted to elucidate the mechanisms of key genes involved in platinum resistance in GC.
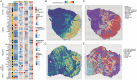
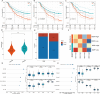
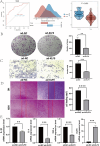
Results: Stomach adenocarcinoma (STAD) patients were classified into three subtypes using the SNF algorithm and multiomics data. Patients with subtype CS2 exhibited a significantly poorer prognosis than those with subtypes CS1 and CS3 (p < 0.05). Subtype CS1 was characterized as immune-deprived, CS2 as stroma-enriched, and CS3 as immune-enriched. Patients with subtype CS2 also exhibited the most adverse therapeutic responses to docetaxel, cisplatin, and gemcitabine. Single-cell analysis revealed high enrichment of M1 module cells with elevated expression of resistance genes, including the transcription factor KLF9. Spatial transcriptomic analysis further confirmed the independent spatial distribution of malignant cells with high expression of drug resistance genes (DRGs). Predictive models based on machine learning demonstrated excellent prognostic performance. Patients in the high DRG group also exhibited poorer responses to immunotherapy. Cellular experiments revealed that KLF9 overexpression significantly inhibited the proliferation of AGS cells (p < 0.05), reduced their resistance to platinum-based drugs, and markedly decreased the levels of inflammatory cytokines in them.
Conclusion: KLF9 was identified as a promising therapeutic target for overcoming platinum resistance in GC, warranting further investigation into its role and potential clinical applications.
Keywords: KLF9; Gastric cancer; Platinum resistance; Similarity network fusion; Spatial transcriptomics.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: Not applicable. Consent for publication: Not applicable. Competing interests: The authors declare no conflict of interest.
Figures









Similar articles
-
Integrating multiomics analysis and machine learning to refine the molecular subtyping and prognostic analysis of stomach adenocarcinoma.Sci Rep. 2025 Jan 30;15(1):3843. doi: 10.1038/s41598-025-87444-3. Sci Rep. 2025. PMID: 39885324 Free PMC article.
-
Integrative single-cell and spatial transcriptomics uncover ELK4-mediated mechanisms in NDUFAB1+ tumor cells driving gastric cancer progression, metabolic reprogramming, and immune evasion.Front Immunol. 2025 Jul 4;16:1591123. doi: 10.3389/fimmu.2025.1591123. eCollection 2025. Front Immunol. 2025. PMID: 40688093 Free PMC article.
-
Integrated proteomics and transcriptomics analysis reveals key regulatory genes between ER-positive/PR-positive and ER-positive/PR-negative breast cancer.BMC Cancer. 2025 Jul 1;25(1):1048. doi: 10.1186/s12885-025-14451-y. BMC Cancer. 2025. PMID: 40597935 Free PMC article.
-
A rapid and systematic review of the clinical effectiveness and cost-effectiveness of paclitaxel, docetaxel, gemcitabine and vinorelbine in non-small-cell lung cancer.Health Technol Assess. 2001;5(32):1-195. doi: 10.3310/hta5320. Health Technol Assess. 2001. PMID: 12065068
-
Topotecan, pegylated liposomal doxorubicin hydrochloride and paclitaxel for second-line or subsequent treatment of advanced ovarian cancer: a systematic review and economic evaluation.Health Technol Assess. 2006 Mar;10(9):1-132. iii-iv. doi: 10.3310/hta10090. Health Technol Assess. 2006. PMID: 16545208
References
-
- Augustin HG, Koh GY. A systems view of the vascular endothelium in health and disease. Cell. 2024;187:4833–58. - PubMed
-
- Barbie DA, Tamayo P, Boehm JS, Kim SY, Moody SE, Dunn IF, Schinzel AC, Sandy P, Meylan E, Scholl C, Frohling S, Chan EM, Sos ML, Michel K, Mermel C, Silver SJ, Weir BA, Reiling JH, Sheng Q, Gupta PB, Wadlow RC, Le H, Hoersch S, Wittner BS, Ramaswamy S, Livingston DM, Sabatini DM, Meyerson M, Thomas RK, Lander ES, Mesirov JP, Root DE, Gilliland DG, Jacks T, Hahn WC. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature. 2009;462:108–12. - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

