This is a preprint.
Bacterial metabolites induce cell wall remodeling, antifungal resistance, and immune recognition of commensal fungi
- PMID: 40777320
- PMCID: PMC12330762
- DOI: 10.1101/2025.07.26.666966
Bacterial metabolites induce cell wall remodeling, antifungal resistance, and immune recognition of commensal fungi
Abstract
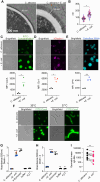
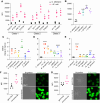
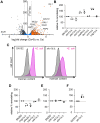
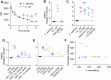
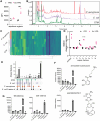
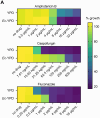
The fungus Candida albicans commensally colonizes mucosal surfaces in healthy individuals but can cause both superficial mucosal and life-threatening disseminated infections. The balance between commensalism and pathogenicity is complex and depends on factors including host and fungal genetic background, the host environment, and fungal interactions with local microbes. The major interaction interface of C. albicans with the host is its multilayered cell wall, which is dynamic and highly responsive to the surrounding environment. Therefore, factors that influence the fungal cell wall will directly impact C. albicans-host interactions. Our work demonstrates that multiple physiologically-relevant gastrointestinal bacteria influence fungal cell wall composition during co-culture with C. albicans, including as complex communities derived from the gut. Using Escherichia coli as a model, we show that bacterial-induced fungal cell wall remodeling occurs rapidly and is mediated by secreted bacterial metabolite(s). Fungal mutant analysis revealed that the high osmolarity glycerol (HOG) pathway, which is critical for responding to environmental stresses, has an important role in regulating this cell wall remodeling phenotype through the Sln1 histidine kinase. Importantly, bacterial-mediated fungal cell wall remodeling increases C. albicans resistance to the echinocandins, increases recognition by both dectin-1 and dectin-2, and decreases recognition by human IgA. Overall, this work comprehensively characterizes an interaction between C. albicans and common gastrointestinal bacteria that has important implications for fungal biology and host interactions.
Figures

Similar articles
-
Histone deacetylase Sir2 promotes the systemic Candida albicans infection by facilitating its immune escape via remodeling the cell wall and maintaining the metabolic activity.mBio. 2024 Jun 12;15(6):e0044524. doi: 10.1128/mbio.00445-24. Epub 2024 Apr 29. mBio. 2024. PMID: 38682948 Free PMC article.
-
Prescription of Controlled Substances: Benefits and Risks.2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 30726003 Free Books & Documents.
-
Commensal colonization of Candida albicans in the mouse gastrointestinal tract is mediated via expression of candidalysin and adhesins.Microbiol Spectr. 2025 Jul 30:e0056725. doi: 10.1128/spectrum.00567-25. Online ahead of print. Microbiol Spectr. 2025. PMID: 40736264
-
Antifungal agents for preventing fungal infections in non-neutropenic critically ill patients.Cochrane Database Syst Rev. 2016 Jan 16;2016(1):CD004920. doi: 10.1002/14651858.CD004920.pub3. Cochrane Database Syst Rev. 2016. PMID: 26772902 Free PMC article.
-
Signs and symptoms to determine if a patient presenting in primary care or hospital outpatient settings has COVID-19.Cochrane Database Syst Rev. 2022 May 20;5(5):CD013665. doi: 10.1002/14651858.CD013665.pub3. Cochrane Database Syst Rev. 2022. PMID: 35593186 Free PMC article.
References
-
- Neville B. A., d’Enfert C. & Bougnoux M.-E. Candida albicans commensalism in the gastrointestinal tract. FEMS Yeast Res. 15, (2015). - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
