This is a preprint.
Mono-mix strategy enables comparative proteomics of a cross-kingdom microbial symbiosis
- PMID: 40777375
- PMCID: PMC12330575
- DOI: 10.1101/2025.07.22.666210
Mono-mix strategy enables comparative proteomics of a cross-kingdom microbial symbiosis
Update in
-
Mono-mix strategy enables comparative proteomics of a cross-kingdom microbial symbiosis.PLoS One. 2026 Jan 16;21(1):e0340253. doi: 10.1371/journal.pone.0340253. eCollection 2026. PLoS One. 2026. PMID: 41544079 Free PMC article.
Abstract
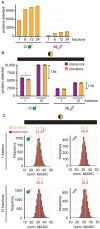
Cross-kingdom microbial symbioses, such as those between algae and bacteria, are key players in biogeochemical cycles. The molecular changes during initiation and establishment of symbiosis are of great interest, but quantitatively monitoring such changes can be challenging, particularly when the microorganisms differ greatly in size or are intimately associated. Here, we analyze output from data-dependent acquisition (DDA) LC-MS/MS proteomics experiments investigating the well-studied interaction between the alga Chlamydomonas reinhardtii and the heterotrophic bacterium Mesorhizobium japonicum. We found that detection of bacterial proteins decreased in coculture by 50% proteome-wide due to the abundance of algal proteins. As a result, standard differential expression analysis led to numerous false-positive reports of significantly downregulated proteins, where it was not possible to distinguish meaningful biological responses to symbiosis from artifacts of the reduced protein detection in coculture relative to monoculture. We show that data normalization alone does not eliminate the impact of altered detection on differential expression analysis of the cross-kingdom symbiosis. We assessed two additional strategies to overcome this methodological artifact inherent to DDA proteomics. In the first, we combined algal and bacterial monocultures at a relative abundance that mimicked the coculture, creating a "mono-mix" control to which the coculture could be compared. This approach enabled comparable detection of bacterial proteins in the coculture and the monoculture control. In the second strategy, we enhanced detection of lowly abundant bacterial proteins by using sample fractionation upstream of LC-MS/MS analysis. When these simple approaches were combined, they allowed for meaningful comparisons of nearly 10,000 algal proteins and over 4,000 bacterial proteins in response to symbiosis by DDA. They successfully recovered expected changes in the bacterial proteome in response to algal coculture, including upregulation of sugar-binding proteins and transporters. They also revealed novel proteomic responses to coculture that guide hypotheses about algal-bacterial interactions.
Keywords: Dual-proteomics; chlorophyte; label-free quantitation; mixed cultures; rhizobia; shotgun proteomics.
Figures




References
-
- Hungate Robert E.. The Rumen and its Microbes. Elsevier; 1966. doi: 10.1016/C2013-0-12555-X - DOI
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
