Phosphoinositide- and Collybistin-Dependent Synaptic Clustering of Gephyrin
- PMID: 40781785
- PMCID: PMC12334862
- DOI: 10.1111/jnc.70169
Phosphoinositide- and Collybistin-Dependent Synaptic Clustering of Gephyrin
Abstract
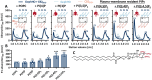
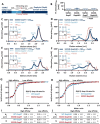
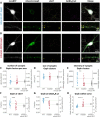
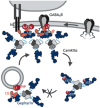
Gephyrin is the main scaffolding protein at inhibitory synapses, clustering glycine and GABAA receptors. At specific GABAergic synapses, the nucleotide exchange factor collybistin recruits gephyrin to the postsynaptic membrane via interaction with phosphoinositides. However, the molecular mechanisms underlying the formation, maintenance, and regulation of collybistin-dependent gephyrin clusters remain poorly understood. This study sheds light on the molecular mechanism of gephyrin cluster formation on the basis of gephyrin self-oligomerization induced by collybistin, leading to the formation of a high-molecular weight (> 5 MDa) gephyrin-collybistin complex, which is regulated in two ways: First, plasma-membrane phosphoinositides promote complex formation, demonstrating their critical role in membrane targeting and stabilization of gephyrin-collybistin clusters at postsynaptic sites. Second, gephyrin phosphorylation at Ser325 abolishes complex formation with collybistin, thus impairing collybistin-dependent gephyrin clustering at GABAergic synapses. Collectively, our data demonstrate a molecular mechanism for synaptic clustering of gephyrin, which involves collybistin- and phosphoinositide-dependent formation of high-molecular weight gephyrin oligomers.
Keywords: GABA receptors; collybistin; gephyrin; inhibitory synapse; phosphoinositides; synaptic clustering.
© 2025 The Author(s). Journal of Neurochemistry published by John Wiley & Sons Ltd on behalf of International Society for Neurochemistry.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures


References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

