The GA4GH Task Execution Application Programming Interface: Enabling Easy Multicloud Task Execution
- PMID: 40792191
- PMCID: PMC12336797
- DOI: 10.1109/mcse.2024.3414994
The GA4GH Task Execution Application Programming Interface: Enabling Easy Multicloud Task Execution
Abstract
The Global Alliance for Genomics and Health (GA4GH) Task Execution Service (TES) application programming interface (API) is a standardized schema and API for describing and executing batch execution tasks. It provides a common way to submit and manage tasks to a variety of compute environments, including on-premises high-performance computing and high-throughput computing systems, cloud computing platforms, and hybrid environments. The TES API is designed to be flexible and extensible, allowing it to be adapted to a wide range of use cases, such as "bringing compute to the data" solutions for federated and distributed data analysis, or load balancing across multicloud infrastructures. This API has been adopted by numerous different service providers and is utilized by several workflow engines, yielding a single abstracted interface for developers and researchers. Using its capabilities, genome research institutes are building extensible hybrid compute systems to study life science.
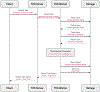
Figures


Similar articles
-
Prescription of Controlled Substances: Benefits and Risks.2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 30726003 Free Books & Documents.
-
Management of urinary stones by experts in stone disease (ESD 2025).Arch Ital Urol Androl. 2025 Jun 30;97(2):14085. doi: 10.4081/aiua.2025.14085. Epub 2025 Jun 30. Arch Ital Urol Androl. 2025. PMID: 40583613 Review.
-
[Volume and health outcomes: evidence from systematic reviews and from evaluation of Italian hospital data].Epidemiol Prev. 2013 Mar-Jun;37(2-3 Suppl 2):1-100. Epidemiol Prev. 2013. PMID: 23851286 Italian.
-
GRAPEVNE - Graphical Analytical Pipeline Development Environment for Infectious Diseases.Wellcome Open Res. 2025 May 27;10:279. doi: 10.12688/wellcomeopenres.23824.1. eCollection 2025. Wellcome Open Res. 2025. PMID: 40585005 Free PMC article.
-
Systemic Inflammatory Response Syndrome.2025 Jun 20. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2025 Jun 20. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 31613449 Free Books & Documents.
References
-
- Voss K, Van der Auwera G, and Gentry J, “Full-stack genomics pipelining with GATK4 + WDL + Cromwell,” F1000Res, vol. 6, Aug. 2017, doi: 10.7490/f1000research.1114634.1. Accessed: Dec. 24, 2023. - DOI
-
- “Openapi-test-runner: An application allowing the TES servers to test their conformance to TES specifications.” GitHub. Accessed: Mar. 22, 2024. [Online]. Available: https://github.com/elixir-cloud-aai/openapi-test-runner
Grants and funding
LinkOut - more resources
Full Text Sources
