METTL3-mediated m6A modification promotes intervertebral disc degeneration
- PMID: 40798940
- PMCID: PMC12351710
- DOI: 10.1080/07853890.2025.2546670
METTL3-mediated m6A modification promotes intervertebral disc degeneration
Abstract
Background: N6-methyladenosine (m6A) modification is a prevalent RNA modification in epigenetics. METTL3, acting as the principal methyltransferase responsible for catalysing m6A, is regarded as a master regulator of this RNA modification. Nonetheless, the complex roles and underlying mechanisms of m6A in relation to intervertebral disc degeneration (IDD) are yet to be fully elucidated. In light of this, this study aimed to explore the intricate functions and mechanisms of METTL3-mediated m6A modification in IDD.
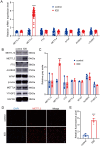
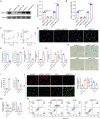
Methods: Our previous batch of RNA sequencing data (GSE167199) and public single-cell data (GSE165722) were utilized to probe the relationship between m6A-related genes and IDD. m6A quantification, RNA m6A immunoblotting, quantitative real-time PCR, western blot and immunofluorescent staining were used to validate the levels of m6A modification and expression of m6A-related genes in nucleus pulposus (NP) tissues and cells. Moreover, gain- and loss-of-function experiments in NP cells were conducted to explore the impact of METTL3 on IDD. In vivo, the effects of METTL3 inhibition and miR-338-3p suppression on IDD progression were assessed.
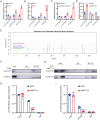
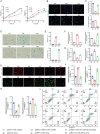
Results: A significant association between METTL3-mediated m6A modification and IDD was identified. Overexpressing METTL3 induced apoptosis, accelerated senescence and inhibited matrix synthesis in NP cells. Additionally, METTL3-mediated m6A modification could expedite the production and maturation of pri-miR-338-3p in NP cells via DGCR8. In vivo, inhibiting METTL3 mitigated IDD progression, while suppressing miR-338-3p notably alleviated IDD during METTL3 overexpression.
Conclusions: This study reveals that targeting METTL3 attenuates IDD progression through the METTL3-m6A-miR-338-3p axis, thereby highlighting the therapeutic potential of METTL3 inhibition for IDD. Future studies should prioritize the development of biomaterial delivery systems for METTL3 inhibitors to ensure both therapeutic protection and sustained, site-specific drug release.
Keywords: Intervertebral disc degeneration; METTL3; N6-methyladenosine; miR-338-3p.
Conflict of interest statement
No potential conflict of interest was reported by the author(s).
Figures


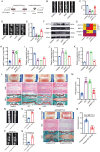
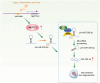

References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
