Comparative Molecular Dynamics Reveals How LRRK2 Inhibitors Distinguish G2019S from Wild-Type
- PMID: 40802138
- PMCID: PMC12350454
- DOI: 10.1007/s11064-025-04520-w
Comparative Molecular Dynamics Reveals How LRRK2 Inhibitors Distinguish G2019S from Wild-Type
Abstract
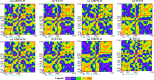
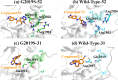
Leucine-rich repeat kinase 2 (LRRK2) has become a critical drug target in Parkinson's disease, with mutation-selective inhibitors offering promising potential for precision medicine. However, the structural similarity between G2019S and wild-type kinases presents a significant challenge in developing selective inhibitors. Although recent advances have led to inhibitors that selectively target G2019S or wild-type kinases, the selectivity mechanism of these inhibitors remains unclear. We employed molecular dynamics simulations to investigate and explore kinase-ligand interactions and identify the underlying mechanisms of selectivity. The results suggest that ligand binding drives the conformational changes, which is a key contributing factor to selectivity, rather than the strength of the ligand binding. The ligand-induced conformational changes lead to kinase destabilisation and inactivation. Additionally, key residues, such as Tyr2018 and Asp2017, were found to play pivotal roles in the selectivity. These insights underscore the importance of incorporating conformational dynamics into the design of future LRRK2 mutant-selective inhibitors.
Keywords: Conformational changes; G2019S/WT selectivity; Kinase-Ligand interaction; LRRK2 inhibitors; Molecular dynamics.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Competing Interests: The authors declare no competing interests. Type: Original Article. Declaration of Competing Interest: All authors have no conflicts of interest to disclose.
Figures
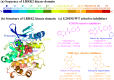


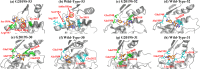

References
-
- Cong C, Milne-Ives M, Ananthakrishnan A, Maetzler W, Meinert E (2025) From past to future: Digital approaches to success of clinical drug trials for Parkinson’s disease. J Parkinson’s Disease:1877718X251330839 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

