Selection Signature Analysis of Whole-Genome Sequences to Identify Genome Differences Between Selected and Unselected Holstein Cattle
- PMID: 40805037
- PMCID: PMC12345450
- DOI: 10.3390/ani15152247
Selection Signature Analysis of Whole-Genome Sequences to Identify Genome Differences Between Selected and Unselected Holstein Cattle
Abstract
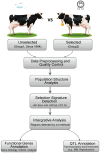
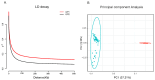
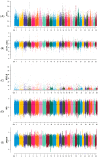
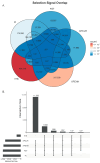
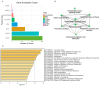
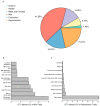
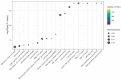
A unique line of Holstein cattle has been maintained without selection in Minnesota since 1964. After many generations, unselected cattle produce less milk, but have better reproductive performance and health traits when compared with contemporary cows. Comparisons between this line of unselected Holstein and those under selection provide useful insights that connect selection and complex traits in cattle. Utilizing these unique resources and sequence data, we sought to identify genome changes due to selection. We sequenced 30 unselected and 54 selected Holstein cattle and compared their sequence variants to identify selection signatures. After many years, the two populations showed completely different patterns in their genome-level population structures and linkage disequilibrium. By integrating signals from five different detection methods, we detected consensus selection signatures from at least four methods covering 14,533 SNPs and 155 protein-coding genes. An integrated analysis of selection signatures with gene annotation, pathways, and the cattle QTL database demonstrated that the genomic regions under selection are related to milk productivity, health, and reproductive efficiency. The polygenic nature of these complex traits is evident from hundreds of selection signatures and candidate genes, suggesting that long-term artificial selection has acted on the whole genome rather than a few major genes. In summary, our study identified candidate selection signatures underlying phenotypic differences between unselected and selected Holstein cows and revealed insights into the genetic basis of complex traits in cattle.
Keywords: Holstein cattle; genome sequence; selection; selection signature.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
Similar articles
-
Prescription of Controlled Substances: Benefits and Risks.2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 30726003 Free Books & Documents.
-
Sequence-based GWAS reveals genes and variants associated with predicted methane emissions in French dairy cows.Genet Sel Evol. 2025 Jun 17;57(1):32. doi: 10.1186/s12711-025-00977-z. Genet Sel Evol. 2025. PMID: 40528185 Free PMC article.
-
Short-Term Memory Impairment.2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 31424720 Free Books & Documents.
-
Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis.Cochrane Database Syst Rev. 2021 Apr 19;4(4):CD011535. doi: 10.1002/14651858.CD011535.pub4. Cochrane Database Syst Rev. 2021. Update in: Cochrane Database Syst Rev. 2022 May 23;5:CD011535. doi: 10.1002/14651858.CD011535.pub5. PMID: 33871055 Free PMC article. Updated.
-
Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis.Cochrane Database Syst Rev. 2020 Jan 9;1(1):CD011535. doi: 10.1002/14651858.CD011535.pub3. Cochrane Database Syst Rev. 2020. Update in: Cochrane Database Syst Rev. 2021 Apr 19;4:CD011535. doi: 10.1002/14651858.CD011535.pub4. PMID: 31917873 Free PMC article. Updated.
References
Grants and funding
LinkOut - more resources
Full Text Sources

