Understanding the Molecular Basis of Miller-Dieker Syndrome
- PMID: 40806509
- PMCID: PMC12347932
- DOI: 10.3390/ijms26157375
Understanding the Molecular Basis of Miller-Dieker Syndrome
Abstract
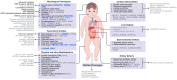
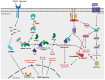
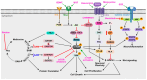
Miller-Dieker Syndrome (MDS) is a rare neurodevelopmental disorder caused by a heterozygous deletion of approximately 26 genes within the MDS locus of human chromosome 17. MDS, which affects 1 in 100,000 babies, can lead to a range of phenotypes, including lissencephaly, severe neurological defects, distinctive facial abnormalities, cognitive impairments, seizures, growth retardation, and congenital heart and liver abnormalities. One hallmark feature of MDS is an unusually smooth brain surface due to abnormal neuronal migration during early brain development. Several genes located within the MDS locus have been implicated in the pathogenesis of MDS, including PAFAH1B1, YWHAE, CRK, and METTL16. These genes play a role in the molecular and cellular pathways that are vital for neuronal migration, the proper development of the cerebral cortex, and protein translation in MDS. Improved model systems, such as MDS patient-derived organoids and multi-omics analyses indicate that WNT/β-catenin signaling, calcium signaling, S-adenosyl methionine (SAM) homeostasis, mammalian target of rapamycin (mTOR) signaling, Janus kinase/signal transducer and activator of transcription (JAK/STAT) signaling, and others are dysfunctional in MDS. This review of MDS integrates details at the clinical level alongside newly emerging details at the molecular and cellular levels, which may inform the development of novel therapeutic strategies for MDS.
Keywords: MDS; gene expression; lissencephaly; rare disease; therapeutics.
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
Prescription of Controlled Substances: Benefits and Risks.2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 30726003 Free Books & Documents.
-
Short-Term Memory Impairment.2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 31424720 Free Books & Documents.
-
Beckwith-Wiedemann Syndrome.2000 Mar 3 [updated 2023 Sep 21]. In: Adam MP, Feldman J, Mirzaa GM, Pagon RA, Wallace SE, Amemiya A, editors. GeneReviews® [Internet]. Seattle (WA): University of Washington, Seattle; 1993–2025. 2000 Mar 3 [updated 2023 Sep 21]. In: Adam MP, Feldman J, Mirzaa GM, Pagon RA, Wallace SE, Amemiya A, editors. GeneReviews® [Internet]. Seattle (WA): University of Washington, Seattle; 1993–2025. PMID: 20301568 Free Books & Documents. Review.
-
Idiopathic (Genetic) Generalized Epilepsy.2024 Feb 12. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2024 Feb 12. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 31536218 Free Books & Documents.
-
The Black Book of Psychotropic Dosing and Monitoring.Psychopharmacol Bull. 2024 Jul 8;54(3):8-59. Psychopharmacol Bull. 2024. PMID: 38993656 Free PMC article. Review.
References
-
- Bershteyn M., Nowakowski T.J., Pollen A.A., Di Lullo E., Nene A., Wynshaw-Boris A., Kriegstein A.R. Human iPSC-Derived Cerebral Organoids Model Cellular Features of Lissencephaly and Reveal Prolonged Mitosis of Outer Radial Glia. Cell Stem Cell. 2017;20:435–449.e4. doi: 10.1016/j.stem.2016.12.007. - DOI - PMC - PubMed
-
- Lindborg B.A., Brekke J.H., Vegoe A.L., Ulrich C.B., Haider K.T., Subramaniam S., Venhuizen S.L., Eide C.R., Orchard P.J., Chen W., et al. Rapid Induction of Cerebral Organoids From Human Induced Pluripotent Stem Cells Using a Chemically Defined Hydrogel and Defined Cell Culture Medium. Stem Cells Transl. Med. 2016;5:970–979. doi: 10.5966/sctm.2015-0305. - DOI - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

