A Natural Language Processing Method Identifies an Association Between Bacterial Communities in the Upper Genital Tract and Ovarian Cancer
- PMID: 40806562
- PMCID: PMC12347966
- DOI: 10.3390/ijms26157432
A Natural Language Processing Method Identifies an Association Between Bacterial Communities in the Upper Genital Tract and Ovarian Cancer
Abstract
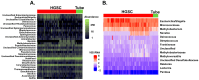
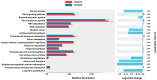
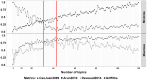
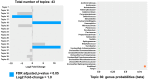
Bacterial communities within the female upper genital tract may influence the risk of ovarian cancer. In this retrospective cohort pilot study, we aim to detect different communities of bacteria between ovarian cancer and normal controls using topic modeling, a natural language processing tool. RNA was extracted and analyzed using the VITCOMIC2 pipeline. Topic modeling assessed differences in bacterial communities. Idatuning identified an optimal latent topic number and Latent Dirichlet Allocation (LDA) assessed topic differences between high-grade serous ovarian cancer (HGSOC) and controls. Results were validated using The Cancer Genome Atlas (TCGA) HGSOC dataset. A total of 801 unique taxa were identified, with 13 bacteria significantly differing between HGSOC and normal controls. LDA modeling revealed a latent topic associated with HGSOC samples, containing bacteria Escherichia/Shigella and Corynebacterineae. Pathway analysis using KEGG databases suggest differences in several biologic pathways including oocyte meiosis, aldosterone-regulated sodium reabsorption, gastric acid secretion, and long-term potentiation. These findings support the hypothesis that bacterial communities in the upper female genital tract may influence the development of HGSOC by altering the local environment, with potential functional implications between HGSOC and normal controls. However, further validation is required to confirms these associations and determine mechanistic relevance.
Keywords: RNA sequencing; RNAseq; microbiome; natural language processing; ovarian cancer; prediction model.
Conflict of interest statement
All the authors have nothing to disclose. This does not alter our adherence to the journal policies on sharing data and materials.
Figures



References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

