Oncodrive3D: fast and accurate detection of structural clusters of somatic mutations under positive selection
- PMID: 40808301
- PMCID: PMC12350100
- DOI: 10.1093/nar/gkaf776
Oncodrive3D: fast and accurate detection of structural clusters of somatic mutations under positive selection
Abstract
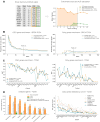
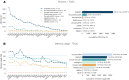
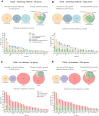
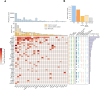
Identifying the genes capable of driving tumorigenesis in different tissues is one of the central goals of cancer genomics. Computational methods that exploit different signals of positive selection in the pattern of mutations observed in genes across tumors have been developed to this end. One such signal of positive selection is clustering of mutations in areas of the three-dimensional (3D) structure of the protein above the expectation under neutrality. Methods that exploit this signal have been hindered by the paucity of proteins with experimentally solved 3D structures covering their entire sequence. Here, we present Oncodrive3D, a computational method that, by exploiting AlphaFold 2 structural models, extends the identification of proteins with significant mutational 3D clusters to the entire human proteome. Oncodrive3D shows sensitivity and specificity on par with state-of-the-art cancer driver gene identification methods exploiting mutational clustering and clearly outperforms them in computational efficiency. We demonstrate, through several examples, how significant mutational 3D clusters identified by Oncodrive3D in different known or potential cancer driver genes can reveal details about the mechanism of tumorigenesis in different cancer types and in clonal hematopoiesis.
© The Author(s) 2025. Published by Oxford University Press.
Conflict of interest statement
None declared.
Figures



References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

