Substituted 1,4-naphthoquinones for potential anticancer therapeutics: In vitro cytotoxic effects and QSAR-guided design of new analogs
- PMID: 40808802
- PMCID: PMC12345971
- DOI: 10.1016/j.csbj.2025.07.040
Substituted 1,4-naphthoquinones for potential anticancer therapeutics: In vitro cytotoxic effects and QSAR-guided design of new analogs
Abstract
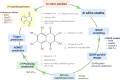
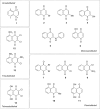
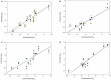
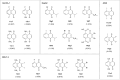
1,4-Naphthoquinone is a promising pharmacophore in drug discovery due to its unique redox reactive nature and wide-ranging bioactivities. Herein, a series of 1,4-naphthoquinones (1-14) were investigated for their anticancer activities against 4 cancer cell lines (i.e., HepG2, HuCCA-1, A549, and MOLT-3). Compound 11 was found to be the most potent and selective anticancer agent against all tested cell lines (IC50 = 0.15 - 1.55 μM, selectivity index = 4.14 - 43.57). QSAR modelling was performed to elucidate key structural features influencing activities against four cancer cell lines. Four QSAR models were successfully constructed using multiple linear regression (MLR) algorithm providing good predictive performance (R: training set = 0.8928-0.9664; testing set = 0.7824-0.9157; RMSE: training set = 0.1755-0.2600; testing set = 0.2726-0.3748). QSAR models suggested that the potent anticancer activities of these naphthoquinones were mainly influenced by polarizability (MATS3p and BELp8), van der Waals volume (GATS5v, GATS6v, and Mor16v), mass (G1m), electronegativity (E1e), and dipole moment (Dipole and EEig15d) as well as ring complexity (RCI) and shape of the compound (SHP2). The models were further applied for guiding the design and predicting activities of an additional set of 248 structurally modified compounds in which the ones with promising predicted activities were highlighted for potential further development. Additionally, pharmacokinetic profiles and possible binding modes towards potential biological targets of the compounds were virtually assessed. Structure-activity relationship analysis was also conducted to highlight key structural features beneficial for further successful design of the related naphthoquinones.
Keywords: ADMET; Anticancer; Computer-aided drug design; Naphthoquinone; QSAR.
© 2025 The Authors.
Conflict of interest statement
The authors declare that they have no conflicts of interest.
Figures



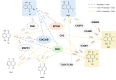

References
-
- Siegel R.L., Giaquinto A.N., Jemal A. Cancer statistics, 2024. CA Cancer J Clin. 2024;74(1):12–49. - PubMed
-
- Bray F., Laversanne M., Sung H., Ferlay J., Siegel R.L., Soerjomataram I., et al. Global cancer statistics 2022: Globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2024;74(3):229–263. - PubMed
LinkOut - more resources
Full Text Sources
