Natrarchaeobius versutus sp. nov. and Natrarchaeobius oligotrophus sp. nov., chitinotrophic natronoarchaea from hypersaline soda lakes, and functional genome analysis of the Natrarchaeobius species
- PMID: 40809049
- PMCID: PMC12343721
- DOI: 10.3389/fmicb.2025.1640521
Natrarchaeobius versutus sp. nov. and Natrarchaeobius oligotrophus sp. nov., chitinotrophic natronoarchaea from hypersaline soda lakes, and functional genome analysis of the Natrarchaeobius species
Abstract
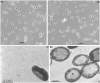
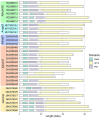
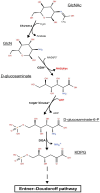
Polysaccharide-degrading natronoarchaea have been poorly studied to date. However, over the past decade, significant progress has been made in understanding their diversity and metabolic potential. In this study, two natronoarchaeal strains, enriched from oxic sediment samples of the soda lakes of Wadi an Natrun in Egypt (AArcel7) and Kulunda steppe in Russia (A-rgal3), were characterized. Strain AArcel7 was enriched with amorphous cellulose, while strain A-rgal3 dominated an enrichment culture using rhamnogalacturonan. Cells of both strains are polymorphic, from motile flat rods to nonmotile cocci. They are aerobic heterotrophs that are able to grow on chitin and several other carbohydrates. Both strains thrive within a salinity range of 2.5 to 4.5 M total Na+, with optimal growth at 3.5–4 M, and are moderately alkaliphilic with an optimum pH at 8.5–9.0 (AArcel7) and 9.2–9.5 (A-rgal3). Genome-based phylogenetic analysis demonstrated that these isolates form a new species lineage in the chitin-specialized genus Natrarchaeobius. An in-depth study of Natrarchaeobius genomes allowed us to identify several genes that potentially enable them to hydrolyze chitin and to metabolize N-acetylglucosamine (GlcNAc), which has not been investigated previously in the chitin-utilizing natronoarchaea. Based on physiological, phylogenetic, and genomic analyses, strains AArcel7 and A-rgal3 are suggested to form a novel species, Natrarchaeobius versutus sp. nov., with AArcel7T (DSM 119357 = UNIQEM U973) as the type strain. Furthermore, strain AArcht7T, formerly classified as the type species of the genus Natrarchaeobius, is proposed to be reclassified as Natrarchaeobius oligotrophus (DSM 119677 = UNIQEM U967).
Keywords: Natrarchaeobius; chitin; chitinase; glycoside hydrolases; natronoarchaea; soda lakes.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
Natrarchaeobius chitinivorans gen. nov., sp. nov., and Natrarchaeobius halalkaliphilus sp. nov., alkaliphilic, chitin-utilizing haloarchaea from hypersaline alkaline lakes.Syst Appl Microbiol. 2019 May;42(3):309-318. doi: 10.1016/j.syapm.2019.01.001. Epub 2019 Jan 8. Syst Appl Microbiol. 2019. PMID: 30638904 Free PMC article.
-
Cellulosispirillum alkaliphilum, gen. nov., sp. nov., an obligately anaerobic cellulotrophic member of the phylum Fibrobacterota from soda lakes.Syst Appl Microbiol. 2025 Jul;48(4):126623. doi: 10.1016/j.syapm.2025.126623. Epub 2025 Jun 2. Syst Appl Microbiol. 2025. PMID: 40482426
-
Corrigendum to "Natrachaeobius chitinivorans gen. nov., sp. nov., and Natrarchaeobius haloalkaliphilus sp. nov., alkaliphilic, chitin-utilizing haloarchaea from hypersaline alkaline lakes" [Syst. Appl. Microbiol. 42 (2019) 309-318].Syst Appl Microbiol. 2019 Sep;42(5):126003. doi: 10.1016/j.syapm.2019.126003. Epub 2019 Aug 2. Syst Appl Microbiol. 2019. PMID: 31378430 Free PMC article. No abstract available.
-
Natronocalculus amylovorans gen. nov., sp. nov., and Natranaeroarchaeum aerophilus sp. nov., dominant culturable amylolytic natronoarchaea from hypersaline soda lakes in southwestern Siberia.Syst Appl Microbiol. 2022 Jul;45(4):126336. doi: 10.1016/j.syapm.2022.126336. Epub 2022 May 20. Syst Appl Microbiol. 2022. PMID: 35644061
-
Where the Little Ones Play the Main Role-Picophytoplankton Predominance in the Soda and Hypersaline Lakes of the Carpathian Basin.Microorganisms. 2022 Apr 14;10(4):818. doi: 10.3390/microorganisms10040818. Microorganisms. 2022. PMID: 35456867 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources

