A pan-genotypic indirect competitive ELISA for serological detection of pigeon circovirus antibodies
- PMID: 40809059
- PMCID: PMC12343533
- DOI: 10.3389/fmicb.2025.1612715
A pan-genotypic indirect competitive ELISA for serological detection of pigeon circovirus antibodies
Abstract
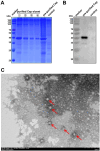
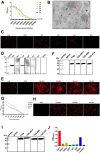
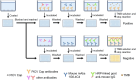
Pigeon circovirus (PiCV) infection, which causes young pigeon disease syndrome (YPDS) and immunosuppression, significantly impacts both the meat and racing pigeon industries. Currently, no inactivated vaccine exists for PiCV prevention, primarily due to the challenges associated with isolating the PiCV virion, except for some gene subunit vaccines express the Cap protein of PiCV. The development of detection techniques is crucial for the diagnosis of PiCV. This study aimed to develop and validate a specific, sensitive indirect competitive enzyme-linked immunosorbent assay (icELISA) for detecting PiCV antibodies in pigeons. We identified the cap gene from a group C PiCV strain (PiCV/Shaanxi/China/10/2021, SX10) isolated from racing pigeons. The Cap of SX10, an immunogen, can self-assemble into virus-like particles (VLPs). A mouse monoclonal antibody (mAb) against Cap, 1G6-4C4, was selected to establish an icELISA. This mAb could identify the PiCV Cap of the strains in groups A to E. The pan-genotypic reactivity of mAb 1G6-4C4 might target a conserved conformational epitope, overcoming limitations of PCR and prior serological assays. The icELISA method exhibited no cross-reactivity with antibodies against other common pigeon pathogens, such as pigeon paramyxovirus type 1 (PPMV-1), avian influenza (H9N2), avian adenovirus type 4 (FAdV-4) or rotavirus (RV). Compared with indirect ELISA (iELISA), icELISA demonstrated comparable performance, as testing of 29 clinical serum samples revealed antibody-positive rates of 51.72% (icELISA) and 44.82% (iELISA), with a 93.10% concordance rate. To a certain extent, icELISA has demonstrated good specificity and sensitivity for detecting PiCV-specific antibodies in pigeons. The developed icELISA provides a robust, specific, and sensitive tool for the serological detection of PiCV infection. Complementary to PCR test, icELISA enhances the comprehensive detection of PICV in epidemiological studies by offering a more practical and sensitive alternative for field applications. Its utility for large-scale epidemiological surveillance in PiCV-endemic regions is validated, highlighting its potential to inform targeted biosecurity and control interventions.
Keywords: Cap; indirect competitive ELISA; monoclonal antibody; pigeon circovirus; virus-like particles.
Copyright © 2025 Wang, Xiao, Zhang, Liu, Tian, Chang, Li, Zhang, Zhang, Li, Yuan and Wang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
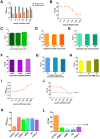
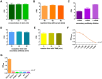
Similar articles
-
Investigating pigeon circovirus infection in a pigeon farm: molecular detection, phylogenetic analysis and complete genome analysis.BMC Genomics. 2024 Apr 16;25(1):369. doi: 10.1186/s12864-024-10303-4. BMC Genomics. 2024. PMID: 38622517 Free PMC article.
-
Prescription of Controlled Substances: Benefits and Risks.2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2025 Jul 6. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 30726003 Free Books & Documents.
-
Antibody tests for identification of current and past infection with SARS-CoV-2.Cochrane Database Syst Rev. 2022 Nov 17;11(11):CD013652. doi: 10.1002/14651858.CD013652.pub2. Cochrane Database Syst Rev. 2022. PMID: 36394900 Free PMC article.
-
Systemic treatments for metastatic cutaneous melanoma.Cochrane Database Syst Rev. 2018 Feb 6;2(2):CD011123. doi: 10.1002/14651858.CD011123.pub2. Cochrane Database Syst Rev. 2018. PMID: 29405038 Free PMC article.
-
Development and validation of dengue virus envelope protein domain III IgG antibody enzyme-linked immunosorbent assay.J Immunol Methods. 2025 Aug;542:113887. doi: 10.1016/j.jim.2025.113887. Epub 2025 May 28. J Immunol Methods. 2025. PMID: 40447073
References
LinkOut - more resources
Full Text Sources
Miscellaneous

